Sanglifehrin-Cyclophilin Interaction: Degradation Work, Synthetic Macrocyclic Analogues, X-ray Crystal Structure and Binding Data
Sedrani, R., Kallen, J., Cabrejas, L.M.M., Papageorgiou, C.D., Senia, F., Rohrbach, S., Wagner, D., Thai, B., Eme, A.-M.J., France, J., Oberer, L., Rihs, G., Zenke, G., Wagner, J.(2003) J Am Chem Soc 125: 3849-3859
- PubMed: 12656618
- DOI: https://doi.org/10.1021/ja021327y
- Primary Citation of Related Structures:
1NMK - PubMed Abstract:
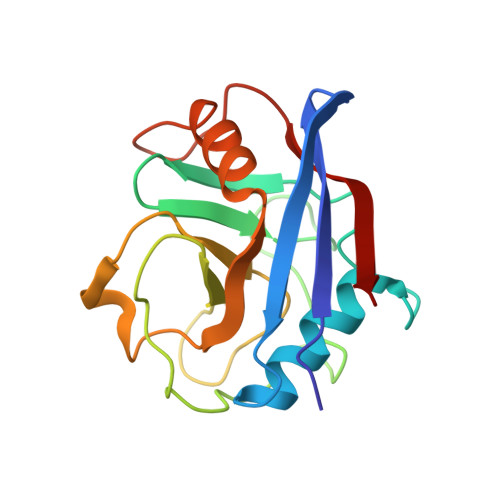
Sanglifehrin A (SFA) is a novel immunosuppressive natural product isolated from Streptomyces sp. A92-308110. SFA has a very strong affinity for cyclophilin A (IC(50) = 6.9 +/- 0.9 nM) but is structurally different from cyclosporin A (CsA) and exerts its immunosuppressive activity via a novel mechanism. SFA has a complex molecular structure consisting of a 22-membered macrocycle, bearing in position 23 a nine-carbon tether terminated by a highly substituted spirobicyclic moiety. Selective oxidative cleavage of the C(26)=C(27) exocyclic double bond affords the spirolactam containing fragment 1 and macrolide 2. The affinity of 2 for cyclophilin (IC(50) = 29 +/- 2.1 nM) is essentially identical to SFA, which indicates that the interaction between SFA and cyclophilin A is mediated exclusively by the macrocyclic portion of the molecule. This observation was confirmed by the X-ray crystal structure resolved at 2.1 A of cyclophilin A complexed to macrolide 16, a close analogue of 2. The X-ray crystal structure showed that macrolide 16 binds to the same deep hydrophobic pocket of cyclophilin A as CsA. Additional valuable details of the structure-activity relationship were obtained by two different chemical approaches: (1) degradation work on macrolide 2 or (2) synthesis of a library of macrolide analogues using the ring-closing metathesis reaction as the key step. Altogether, it appears that the complex macrocyclic fragment of SFA is a highly optimized combination of multiple functionalities including an (E,E)-diene, a short polypropionate fragment, and an unusual tripeptide unit, which together provide an extremely strong affinity for cyclophilin A.
Organizational Affiliation:
Transplantation Research, Novartis Pharma AG, S-507.312, CH-4002 Basel, Switzerland.