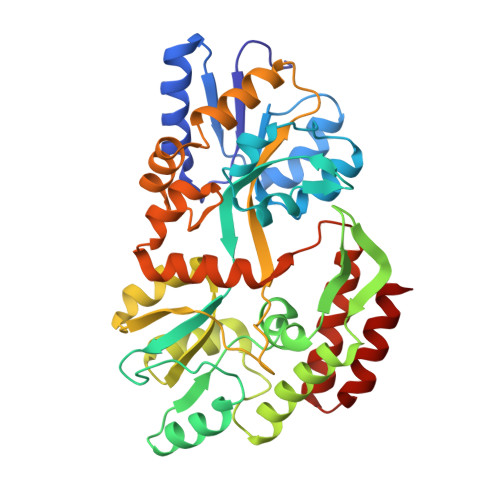
Crystallographic evidence of a large ligand-induced hinge-twist motion between the two domains of the maltodextrin binding protein involved in active transport and chemotaxis.
Sharff, A.J., Rodseth, L.E., Spurlino, J.C., Quiocho, F.A.(1992) Biochemistry 31: 10657-10663
- PubMed: 1420181
- DOI: https://doi.org/10.1021/bi00159a003
- Primary Citation of Related Structures:
1OMP - PubMed Abstract:
The periplasmic maltodextrin binding protein of Escherichia coli serves as an initial receptor for the active transport of and chemotaxis toward maltooligosaccharides. The three-dimensional structure of the binding protein complexed with maltose has been previously reported [Spurlino, J. C., Lu, G.-Y., & Quiocho, F. A. (1991) J. Biol. Chem. 266, 5202-5219]. Here we report the structure of the unliganded form of the binding protein refined to 1.8-A resolution. This structure, combined with that for the liganded form, provides the first crystallographic evidence that a major ligand-induced conformational change occurs in a periplasmic binding protein. The unliganded structure shows a rigid-body "hinge-bending" between the two globular domains by approximately 35 degrees, relative to the maltose-bound structure, opening the sugar binding site groove located between the two domains. In addition, there is an 8 degrees twist of one domain relative to the other domain. The conformational changes observed between this structure and the maltose-bound structure are consistent with current models of maltose/maltodextrin transport and maltose chemotaxis and solidify a mechanism for receptor differentiation between the ligand-free and ligand-bound forms in signal transduction.
Organizational Affiliation:
Howard Hughes Medical Institute, Baylor College of Medicine, Houston, Texas 77030.