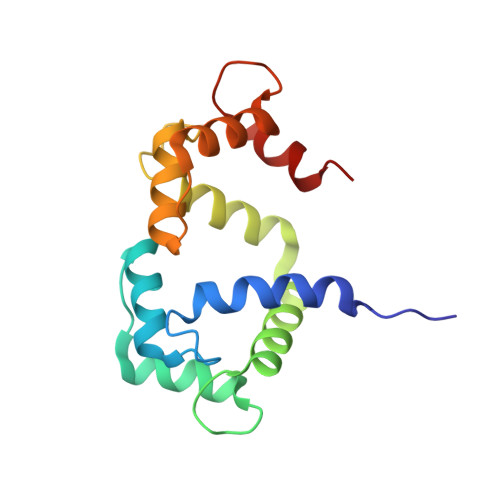
Structure of calmodulin complexed with an olfactory CNG channel fragment and role of the central linker: residual dipolar couplings to evaluate calmodulin binding modes outside the kinase family.
Contessa, G.M., Orsale, M., Melino, S., Torre, V., Paci, M., Desideri, A., Cicero, D.O.(2005) J Biomol NMR 31: 185-199
- PubMed: 15803393
- DOI: https://doi.org/10.1007/s10858-005-0165-1
- Primary Citation of Related Structures:
1SY9 - PubMed Abstract:
The NMR high-resolution structure of calmodulin complexed with a fragment of the olfactory cyclic-nucleotide gated channel is described. This structure shows features that are unique for this complex, including an active role of the linker connecting the N- and C-lobes of calmodulin upon binding of the peptide. Such linker is not only involved in the formation of an hydrophobic pocket to accommodate a bulky peptide residue, but it also provides a positively charged region complementary to a negative charge of the target. This complex of calmodulin with a target not belonging to the kinase family was used to test the residual dipolar coupling (RDC) approach for the determination of calmodulin binding modes to peptides. Although the complex here characterized belongs to the (1--14) family, high Q values were obtained with all the 1:1 complexes for which crystalline structures are available. Reduction of the RDC data set used for the correlation analysis to structured regions of the complex allowed a clear identification of the binding mode. Excluded regions comprise calcium binding loops and loops connecting the EF-hand motifs.
Organizational Affiliation:
Department of Chemical Sciences and Technologies, University of Rome Tor Vergata, via della Ricerca Scientifica, 00133, Rome, Italy.