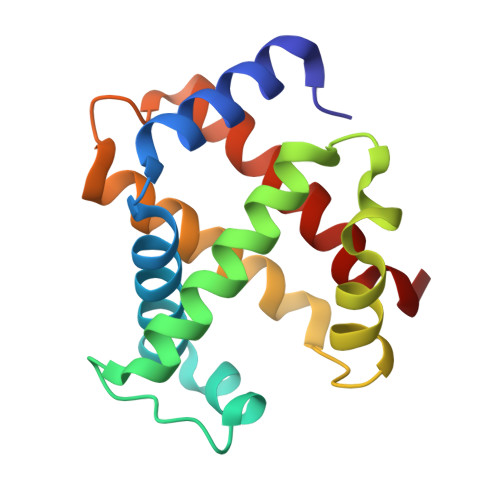
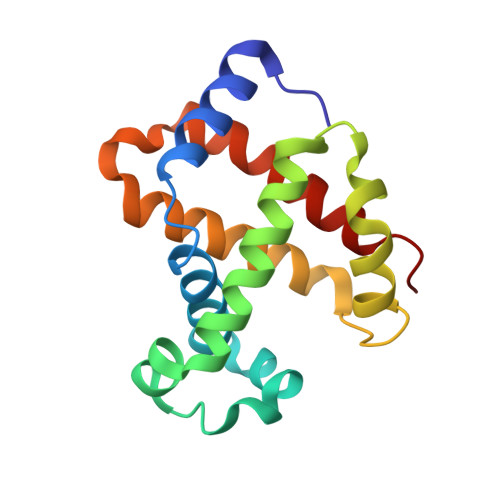
The enigma of the liganded hemoglobin end state: a novel quaternary structure of human carbonmonoxy hemoglobin.
Safo, M.K., Abraham, D.J.(2005) Biochemistry 44: 8347-8359
- PubMed: 15938624
- DOI: https://doi.org/10.1021/bi050412q
- Primary Citation of Related Structures:
1MKO, 1YZI - PubMed Abstract:
The liganded hemoglobin (Hb) high-salt crystallization condition described by Max Perutz has generated three different crystals of human adult carbonmonoxy hemoglobin (COHbA). The first crystal is isomorphous with the "classical" liganded or R Hb structure. The second crystal reveals a new liganded Hb quaternary structure, RR2, that assumes an intermediate conformation between the R form and another liganded Hb quaternary structure, R2, which was discovered more than a decade ago. Like the R2 structure, the diagnostic R state hydrogen bond between beta2His97 and alpha1Thr38 is missing in the RR2 structure. The third crystal adopts a novel liganded Hb conformation, which we have termed R3, and it shows substantial quaternary structural differences from the R, RR2, and R2 structures. The quaternary structure differences between T and R3 are as large as those between T and R2; however, the T --> R3 and T --> R2 transitions are in different directions as defined by rigid-body screw rotation. Moreover, R3 represents an end state. Compared to all known liganded Hb structures, R3 shows remarkably reduced strain at the alpha-heme, reduced steric contact between the beta-heme ligand and the distal residues, smaller alpha- and beta-clefts, and reduced alpha1-alpha2 and beta1-beta2 iron-iron distances. Together, these unique structural features in R3 should make it the most relaxed and/or greatly enhance its affinity for oxygen compared to the other liganded Hbs. The current Hb structure-function relationships that are now based on T --> R, T -->R --> R2, or T --> R2 --> R transitions may have to be reexamined to take into account the RR2 and R3 liganded structures.
Organizational Affiliation:
School of Pharmacy, Department of Medicinal Chemistry, and Institute for Structural Biology and Drug Discovery, Virginia Commonwealth University, Richmond, Virginia 23219, USA. msafo@mail2.vcu.edu