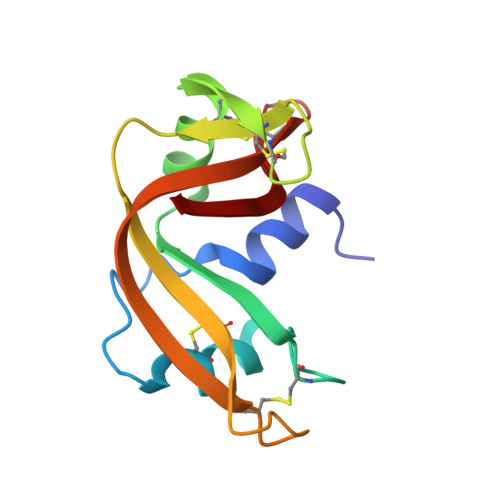
High-resolution three-dimensional structure of ribonuclease A in solution by nuclear magnetic resonance spectroscopy.
Santoro, J., Gonzalez, C., Bruix, M., Neira, J.L., Nieto, J.L., Herranz, J., Rico, M.(1993) J Mol Biol 229: 722-734
- PubMed: 8381876
- DOI: https://doi.org/10.1006/jmbi.1993.1075
- Primary Citation of Related Structures:
2AAS - PubMed Abstract:
High-resolution three-dimensional structures of bovine pancreatic ribonuclease A in aqueous solution have been determined by nuclear magnetic resonance (NMR) spectroscopy combined with restrained molecular dynamics calculations. The structures are based on: (1) 464 interproton distance constraints with accurate upper and lower limits, determined from build-up rates of nuclear Overhauser effects (NOE) by using the complete relaxation matrix; (2) 999 more approximate upper limits for interproton distances; and (3) 42 dihedral angle constraints (37 for phi and 5 for chi 1). A total of 16 structures were calculated, which show a root-mean-square (r.m.s.) deviation of 0.66 A for the backbone atoms and 1.68 A for all heavy-atoms. The converged structures are highly similar to those found in the crystal state. r.m.s. deviation of backbone atom positions in the crystal as compared to those in the average solution structure is 0.92 A. Observed differences are concentrated in loop regions and in the neighborhood of His119 and His48 side-chains. Dynamic aspects, such as H/D amide proton exchange and side-chain mobility have been examined.
Organizational Affiliation:
Instituto de Estructura de la Materia, CSIC, Madrid, Spain.