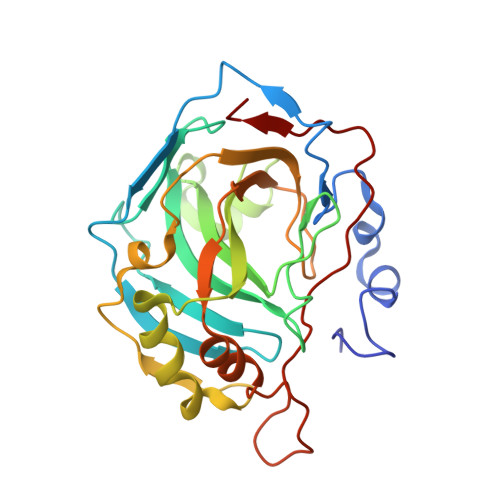
Carbonic anhydrase activators: X-ray crystal structure of the adduct of human isozyme II with l-histidine as a platform for the design of stronger activators.
Temperini, C., Scozzafava, A., Puccetti, L., Supuran, C.T.(2005) Bioorg Med Chem Lett 15: 5136-5141
- PubMed: 16214338
- DOI: https://doi.org/10.1016/j.bmcl.2005.08.069
- Primary Citation of Related Structures:
2ABE - PubMed Abstract:
Activation of the carbonic anhydrase (CA, EC 4.2.1.1) isoforms hCA I, II, and IV with l-histidine and some of its derivatives has been investigated by kinetic and X-ray crystallographic methods. l-His was a potent activator of isozymes I and IV (activation constants in the range of 4-33microM), and a moderate hCA II activator (activation constant of 113microM). Both carboxy- as well as amino-substituted l-His derivatives, such as the methyl ester or the dipeptide carnosine (beta-Ala-His), acted as more efficient activators as compared to l-His. The X-ray crystallographic structure of the hCA II-l-His adduct showed the activator to be anchored at the entrance of the active site cavity, participating in an extended network of hydrogen bonds with the amino acid residues His64, Asn67, and Gln92 and, with three water molecules connecting it to the zinc-bound water. Although the binding site of l-His is similar to that of histamine, the first CA activator for which the X-ray crystal structure has been reported in complex with hCA II (Briganti, F.; Mangani, S.; Orioli, P.; Scozzafava, A.; Vernaglione, G.; Supuran, C. T. Biochemistry1997, 36, 10384) there are important differences of binding between the two structurally related activators, since histamine interacts among others with Asn67 and Gln92 (similarly to l-His), but also with Asn62 and not His64, whereas the number of water molecules connecting them to the zinc-bound water is different (two for histamine, three for l-His). Furthermore, the imidazole moieties of the two activators adopt different conformations when bound to the enzyme active site. Since neither the amino- nor carboxy moieties of l-His participate in interactions with amino acid moieties of the active site, they can be derivatized for obtaining more potent activators, with pharmacological applications for the enhancement of synaptic efficacy. This may constitute a novel approach for the treatment of Alzheimer's disease, aging, and other conditions in need of achieving spatial learning and memory therapy.
Organizational Affiliation:
Università degli Studi di Firenze, Laboratorio di Chimica Bioinorganica, Rm. 188, Via della Lastruccia 3, I-50019 Sesto Fiorentino, Firenze, Italy.