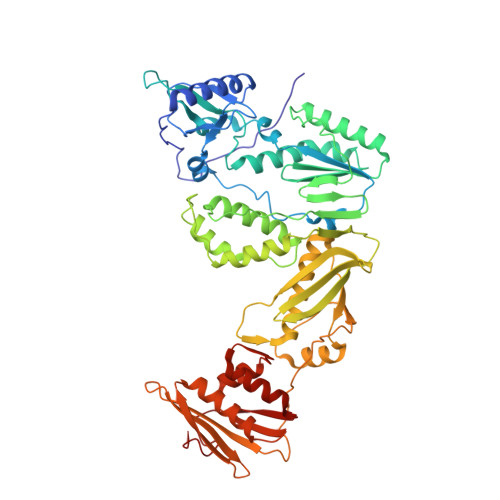
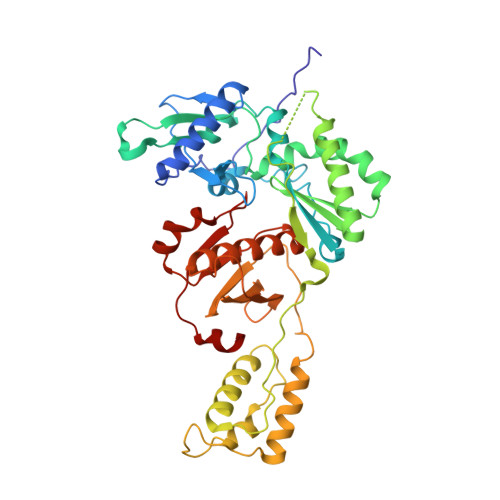
HIV-1 Reverse Transcriptase (RT) Polymorphism 172K Suppresses the Effect of Clinically Relevant Drug Resistance Mutations to Both Nucleoside and Non-nucleoside RT Inhibitors.
Hachiya, A., Marchand, B., Kirby, K.A., Michailidis, E., Tu, X., Palczewski, K., Ong, Y.T., Li, Z., Griffin, D.T., Schuckmann, M.M., Tanuma, J., Oka, S., Singh, K., Kodama, E.N., Sarafianos, S.G.(2012) J Biol Chem 287: 29988-29999
- PubMed: 22761416
- DOI: https://doi.org/10.1074/jbc.M112.351551
- Primary Citation of Related Structures:
4DG1 - PubMed Abstract:
Polymorphisms have poorly understood effects on drug susceptibility and may affect the outcome of HIV treatment. We have discovered that an HIV-1 reverse transcriptase (RT) polymorphism (RT(172K)) is present in clinical samples and in widely used laboratory strains (BH10), and it profoundly affects HIV-1 susceptibility to both nucleoside (NRTIs) and non-nucleoside RT inhibitors (NNRTIs) when combined with certain mutations. Polymorphism 172K significantly suppressed zidovudine resistance caused by excision (e.g. thymidine-associated mutations) and not by discrimination mechanism mutations (e.g. Q151M complex). Moreover, it attenuated resistance to nevirapine or efavirenz imparted by NNRTI mutations. Although 172K favored RT-DNA binding at an excisable pre-translocation conformation, it decreased excision by thymidine-associated mutation-containing RT. 172K affected DNA handling and decreased RT processivity without significantly affecting the k(cat)/K(m) values for dNTP. Surface plasmon resonance experiments revealed that RT(172K) decreased DNA binding by increasing the dissociation rate. Hence, the increased zidovudine susceptibility of RT(172K) results from its increased dissociation from the chain-terminated DNA and reduced primer unblocking. We solved a high resolution (2.15 Å) crystal structure of RT mutated at 172 and compared crystal structures of RT(172R) and RT(172K) bound to NNRTIs or DNA/dNTP. Our structural analyses highlight differences in the interactions between α-helix E (where 172 resides) and the active site β9-strand that involve the YMDD loop and the NNRTI binding pocket. Such changes may increase dissociation of DNA, thus suppressing excision-based NRTI resistance and also offset the effect of NNRTI resistance mutations thereby restoring NNRTI binding.
Organizational Affiliation:
Christopher S. Bond Life Sciences Center, Department of Molecular Microbiology and Immunology, University of Missouri School of Medicine, Columbia, MO 65211, USA.