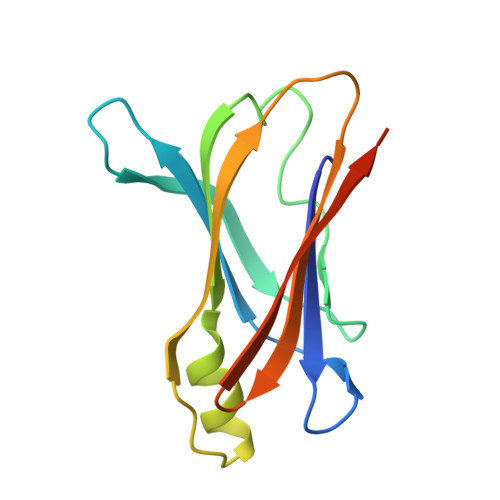
Uncovering the Mechanism of Aggregation of Human Transthyretin.
Saelices, L., Johnson, L.M., Liang, W.Y., Sawaya, M.R., Cascio, D., Ruchala, P., Whitelegge, J., Jiang, L., Riek, R., Eisenberg, D.S.(2015) J Biol Chem 290: 28932-28943
- PubMed: 26459562
- DOI: https://doi.org/10.1074/jbc.M115.659912
- Primary Citation of Related Structures:
4TKW, 4TL4, 4TL5, 4TLK, 4TLS, 4TLT, 4TM9, 4TNE, 4XFN, 4XFO - PubMed Abstract:
The tetrameric thyroxine transport protein transthyretin (TTR) forms amyloid fibrils upon dissociation and monomer unfolding. The aggregation of transthyretin has been reported as the cause of the life-threatening transthyretin amyloidosis. The standard treatment of familial cases of TTR amyloidosis has been liver transplantation. Although aggregation-preventing strategies involving ligands are known, understanding the mechanism of TTR aggregation can lead to additional inhibition approaches. Several models of TTR amyloid fibrils have been proposed, but the segments that drive aggregation of the protein have remained unknown. Here we identify β-strands F and H as necessary for TTR aggregation. Based on the crystal structures of these segments, we designed two non-natural peptide inhibitors that block aggregation. This work provides the first characterization of peptide inhibitors for TTR aggregation, establishing a novel therapeutic strategy.
Organizational Affiliation:
From the Department of Biological Chemistry, Department of Chemistry and Biochemistry, and Howard Hughes Medical Institute, UCLA, Los Angeles, California 90095-1570, Swiss Federal Institute of Technology in Zürich (ETH), Physical Chemistry, ETH Hönggerberg, 8093 Zürich, Switzerland, and.