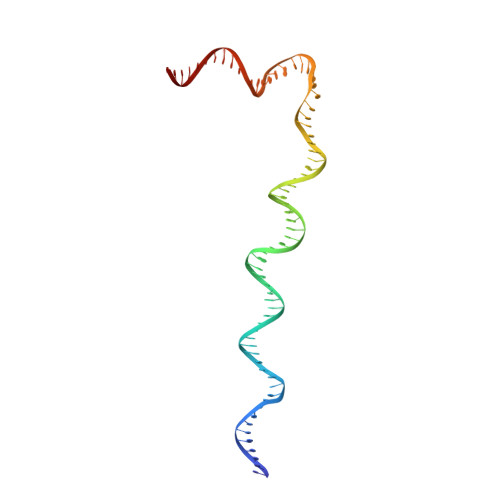
Structure of an RNA Polymerase II Pre-Initiation Complex
Murakami, K., Tsai, K., Kalisman, N., Bushnell, D.A., Asturias, F.J., Kornberg, R.D.(2015) Proc Natl Acad Sci U S A 112: 13543
- PubMed: 26483468
- DOI: https://doi.org/10.1073/pnas.1518255112
- Primary Citation of Related Structures:
5FMF - PubMed Abstract:
The structure of a 33-protein, 1.5-MDa RNA polymerase II preinitiation complex (PIC) was determined by cryo-EM and image processing at a resolution of 6-11 Å. Atomic structures of over 50% of the mass were fitted into the electron density map in a manner consistent with protein-protein cross-links previously identified by mass spectrometry. The resulting model of the PIC confirmed the main conclusions from previous cryo-EM at lower resolution, including the association of promoter DNA only with general transcription factors and not with the polymerase. Electron density due to DNA was identifiable by the grooves of the double helix and exhibited sharp bends at points downstream of the TATA box, with an important consequence: The DNA at the downstream end coincides with the DNA in a transcribing polymerase. The structure of the PIC is therefore conducive to promoter melting, start-site scanning, and the initiation of transcription.
Organizational Affiliation:
Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305; Department of Biochemistry and Biophysics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104;