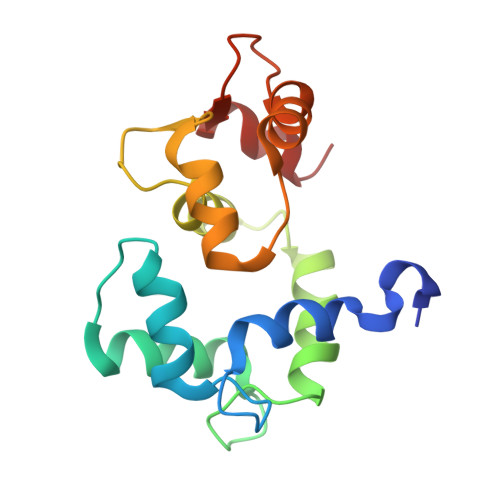
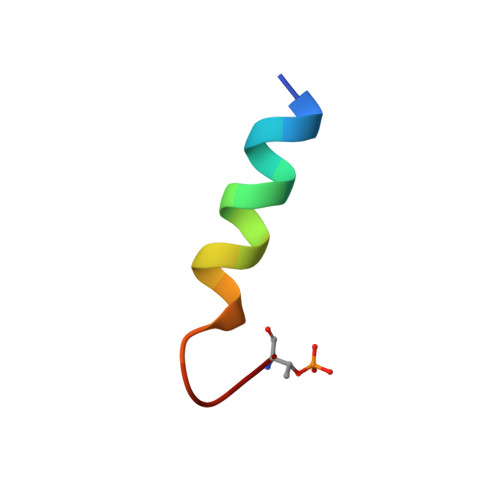
Ca2+/calmodulin binding to PSD-95 mediates homeostatic synaptic scaling down.
Chowdhury, D., Turner, M., Patriarchi, T., Hergarden, A.C., Anderson, D., Zhang, Y., Sun, J., Chen, C.Y., Ames, J.B., Hell, J.W.(2018) EMBO J 37: 122-138
- PubMed: 29118000
- DOI: https://doi.org/10.15252/embj.201695829
- Primary Citation of Related Structures:
5J7J - PubMed Abstract:
Postsynaptic density protein-95 (PSD-95) localizes AMPA-type glutamate receptors (AMPARs) to postsynaptic sites of glutamatergic synapses. Its postsynaptic displacement is necessary for loss of AMPARs during homeostatic scaling down of synapses. Here, we demonstrate that upon Ca 2+ influx, Ca 2+ /calmodulin (Ca 2+ /CaM) binding to the N-terminus of PSD-95 mediates postsynaptic loss of PSD-95 and AMPARs during homeostatic scaling down. Our NMR structural analysis identified E17 within the PSD-95 N-terminus as important for binding to Ca 2+ /CaM by interacting with R126 on CaM. Mutating E17 to R prevented homeostatic scaling down in primary hippocampal neurons, which is rescued via charge inversion by ectopic expression of CaM R 126E , as determined by analysis of miniature excitatory postsynaptic currents. Accordingly, increased binding of Ca 2+ /CaM to PSD-95 induced by a chronic increase in Ca 2+ influx is a critical molecular event in homeostatic downscaling of glutamatergic synaptic transmission.
Organizational Affiliation:
Department of Pharmacology, University of California, Davis, CA, USA.