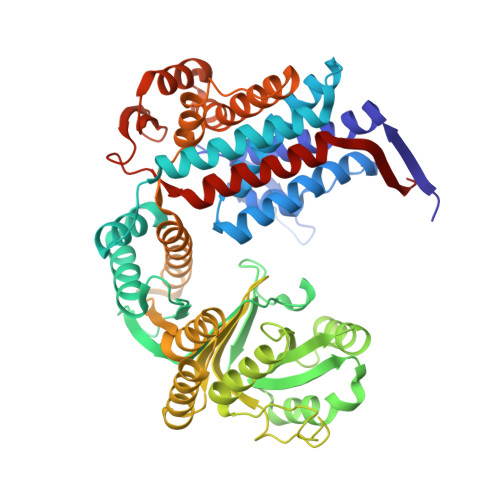
Subunit conformational variation within individual GroEL oligomers resolved by Cryo-EM.
Roh, S.H., Hryc, C.F., Jeong, H.H., Fei, X., Jakana, J., Lorimer, G.H., Chiu, W.(2017) Proc Natl Acad Sci U S A 114: 8259-8264
- PubMed: 28710336
- DOI: https://doi.org/10.1073/pnas.1704725114
- Primary Citation of Related Structures:
5W0S - PubMed Abstract:
Single-particle electron cryo-microscopy (cryo-EM) is an emerging tool for resolving structures of conformationally heterogeneous particles; however, each structure is derived from an average of many particles with presumed identical conformations. We used a 3.5-Å cryo-EM reconstruction with imposed D7 symmetry to further analyze structural heterogeneity among chemically identical subunits in each GroEL oligomer. Focused classification of the 14 subunits in each oligomer revealed three dominant classes of subunit conformations. Each class resembled a distinct GroEL crystal structure in the Protein Data Bank. The conformational differences stem from the orientations of the apical domain. We mapped each conformation class to its subunit locations within each GroEL oligomer in our dataset. The spatial distributions of each conformation class differed among oligomers, and most oligomers contained 10-12 subunits of the three dominant conformation classes. Adjacent subunits were found to more likely assume the same conformation class, suggesting correlation among subunits in the oligomer. This study demonstrates the utility of cryo-EM in revealing structure dynamics within a single protein oligomer.
Organizational Affiliation:
National Center for Macromolecular Imaging, Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, TX 77030.