Broadly Reactive Human Monoclonal Antibodies Elicited following Pandemic H1N1 Influenza Virus Exposure Protect Mice against Highly Pathogenic H5N1 Challenge.
Nachbagauer, R., Shore, D., Yang, H., Johnson, S.K., Gabbard, J.D., Tompkins, S.M., Wrammert, J., Wilson, P.C., Stevens, J., Ahmed, R., Krammer, F., Ellebedy, A.H.(2018) J Virol 92
- PubMed: 29899095
- DOI: https://doi.org/10.1128/JVI.00949-18
- Primary Citation of Related Structures:
6B3M - PubMed Abstract:
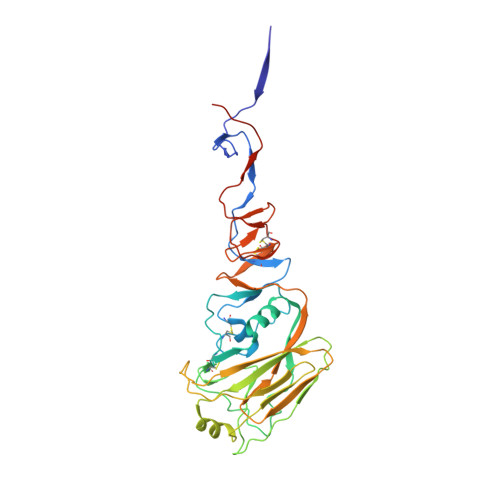
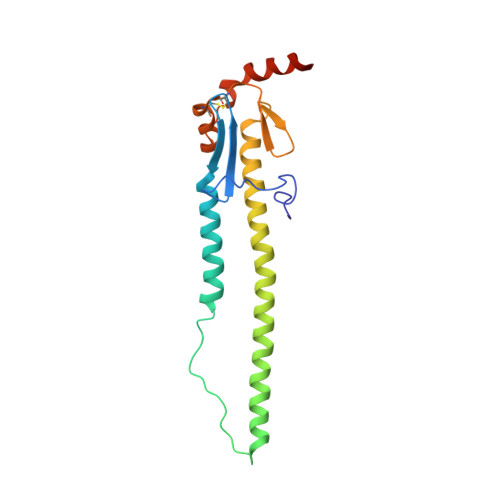
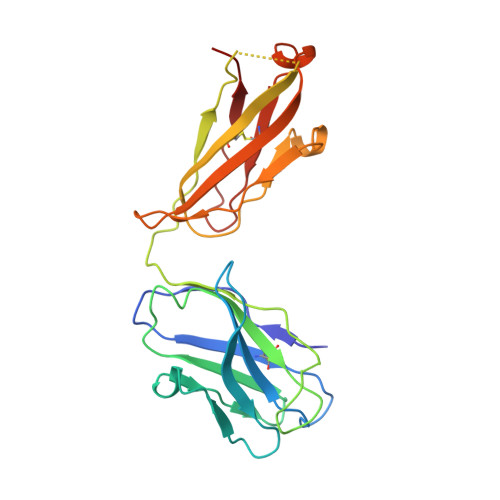
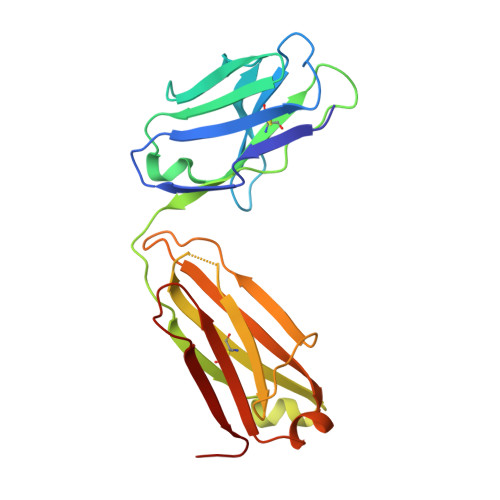
Broadly cross-reactive antibodies (Abs) that recognize conserved epitopes within the influenza virus hemagglutinin (HA) stalk domain are of particular interest for their potential use as therapeutic and prophylactic agents against multiple influenza virus subtypes, including zoonotic virus strains. Here, we characterized four human HA stalk-reactive monoclonal antibodies (MAbs) for their binding breadth and affinity, in vitro neutralization capacity, and in vivo protective potential against an highly pathogenic avian influenza virus. The monoclonal antibodies were isolated from individuals shortly following infection with (70-1F02 and 1009-3B05) or vaccination against (05-2G02 and 09-3A01) A(H1N1)pdm09. Three of the MAbs bound HAs from multiple strains of group 1 viruses, and one MAb, 05-2G02, bound to both group 1 and group 2 influenza A virus HAs. All four antibodies prophylactically protected mice against a lethal challenge with the highly pathogenic A/Vietnam/1203/04 (H5N1) strain. Two MAbs, 70-1F02 and 09-3A01, were further tested for their therapeutic efficacy against the same strain and showed good efficacy in this setting as well. One MAb, 70-1F02, cocrystallized with H5 HA and showed heavy-chain-only interactions similar to those seen with the previously described CR6261 anti-stalk antibody. Finally, we show that antibodies that compete with these MAbs are prevalent in serum from an individual recently infected with the A(H1N1)pdm09 virus. The antibodies described here can be developed into broad-spectrum antiviral therapeutics that could be used to combat infections by zoonotic or emerging pandemic influenza viruses. IMPORTANCE The rise in zoonotic infections of humans by emerging influenza viruses is a worldwide public health concern. The majority of recent zoonotic human influenza cases were caused by H7N9 and H5Nx viruses and were associated with high morbidity and mortality. In addition, seasonal influenza viruses are estimated to cause up to 650,000 deaths annually worldwide. Currently available antiviral treatment options include only neuraminidase inhibitors, but some influenza viruses are naturally resistant to these drugs, and others quickly develop resistance-conferring mutations. Alternative therapeutics are urgently needed. Broadly protective antibodies that target the conserved "stalk" domain of the hemagglutinin represent potential potent antiviral prophylactic and therapeutic agents that can assist pandemic preparedness. Here, we describe four human monoclonal antibodies that target conserved regions of influenza HA and characterize their binding spectrum as well as their protective capacity in prophylactic and therapeutic settings against a lethal challenge with a zoonotic influenza virus.
Organizational Affiliation:
Department of Microbiology, Icahn School of Medicine at Mount Sinai, New York, New York, USA.