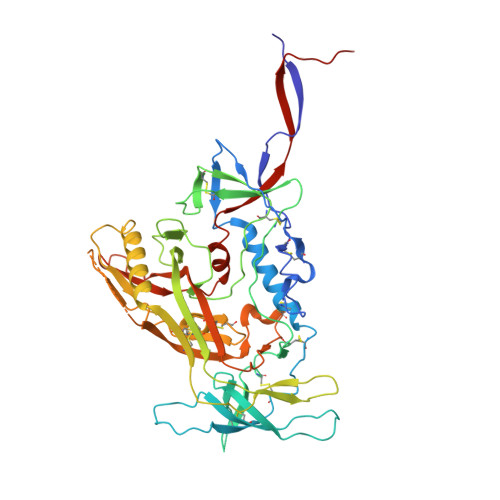
Interdomain Stabilization Impairs CD4 Binding and Improves Immunogenicity of the HIV-1 Envelope Trimer.
Zhang, P., Gorman, J., Geng, H., Liu, Q., Lin, Y., Tsybovsky, Y., Go, E.P., Dey, B., Andine, T., Kwon, A., Patel, M., Gururani, D., Uddin, F., Guzzo, C., Cimbro, R., Miao, H., McKee, K., Chuang, G.Y., Martin, L., Sironi, F., Malnati, M.S., Desaire, H., Berger, E.A., Mascola, J.R., Dolan, M.A., Kwong, P.D., Lusso, P.(2018) Cell Host Microbe 23: 832-844.e6
- PubMed: 29902444
- DOI: https://doi.org/10.1016/j.chom.2018.05.002
- Primary Citation of Related Structures:
6DE7 - PubMed Abstract:
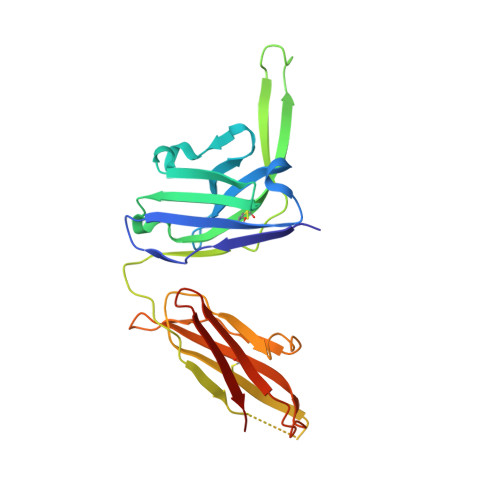
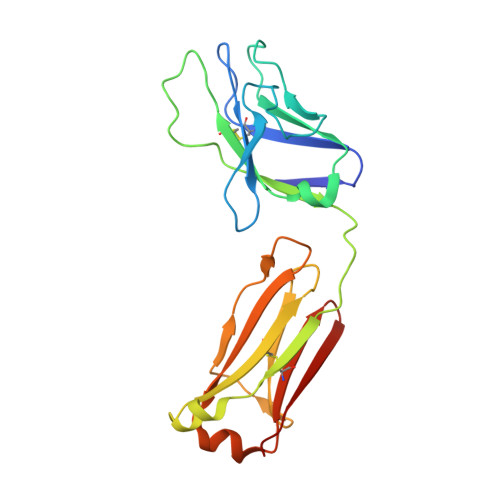
The HIV-1 envelope (Env) spike is a trimer of gp120/gp41 heterodimers that mediates viral entry. Binding to CD4 on the host cell membrane is the first essential step for infection but disrupts the native antigenic state of Env, posing a key obstacle to vaccine development. We locked the HIV-1 Env trimer in a pre-fusion configuration, resulting in impaired CD4 binding and enhanced binding to broadly neutralizing antibodies. This design was achieved via structure-guided introduction of neo-disulfide bonds bridging the gp120 inner and outer domains and was successfully applied to soluble trimers and native gp160 from different HIV-1 clades. Crystallization illustrated the structural basis for CD4-binding impairment. Immunization of rabbits with locked trimers from two different clades elicited neutralizing antibodies against tier-2 viruses with a repaired glycan shield regardless of treatment with a functional CD4 mimic. Thus, interdomain stabilization provides a widely applicable template for the design of Env-based HIV-1 vaccines.
Organizational Affiliation:
Laboratory of Immunoregulation, National Institute of Allergy and Infectious Diseases, NIH, Bethesda, MD 20892, USA.