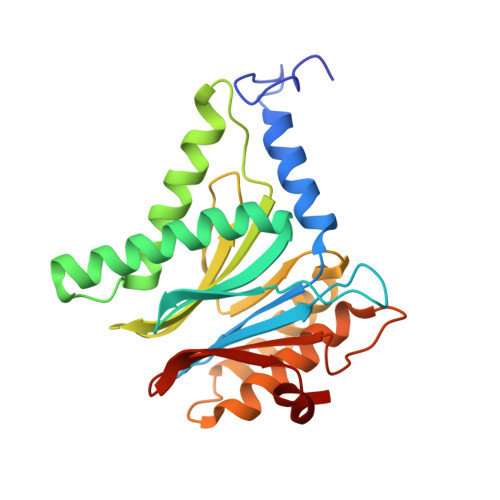
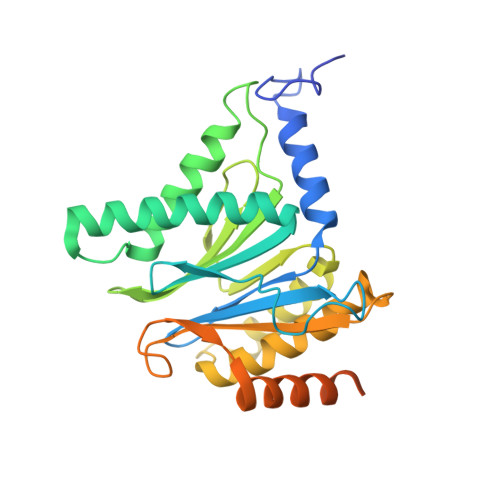
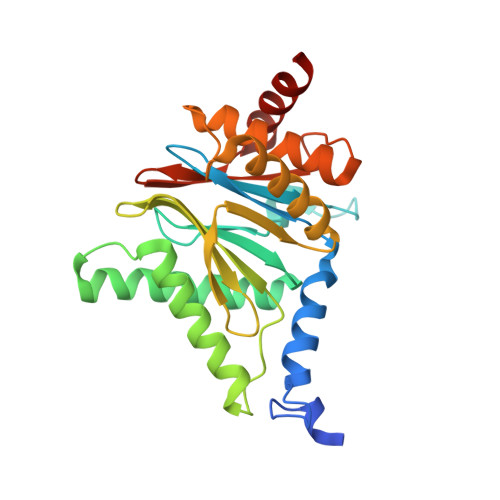
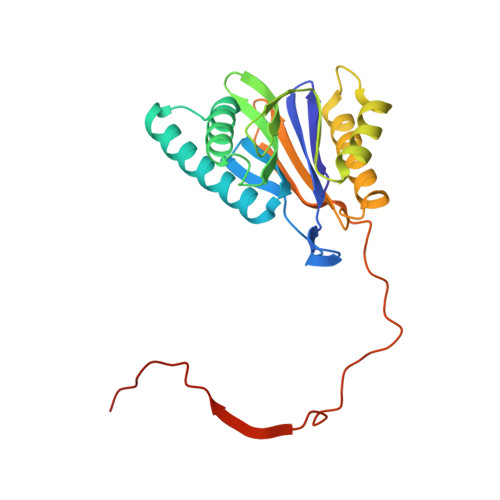
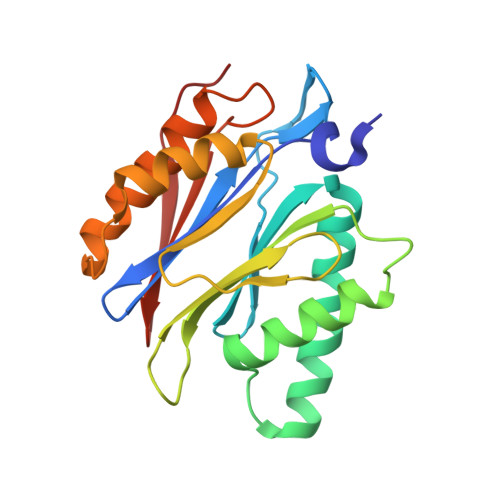
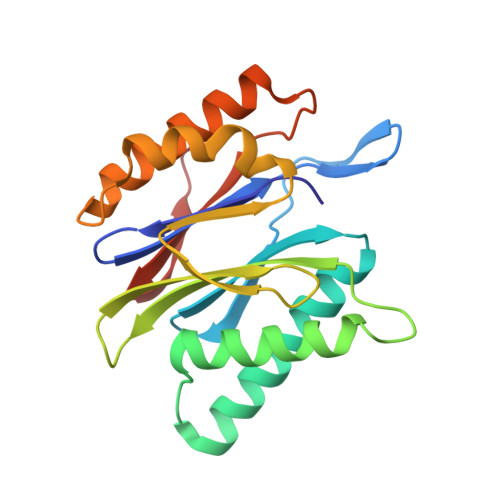
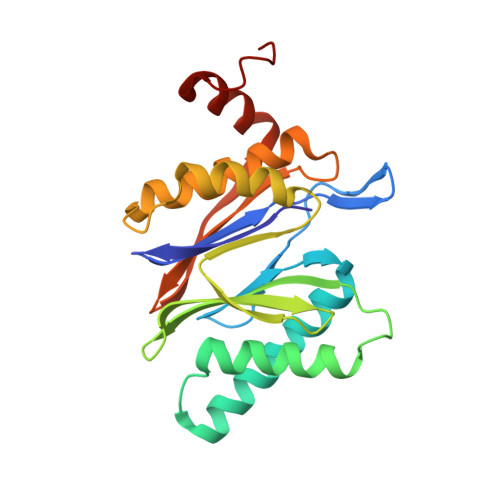
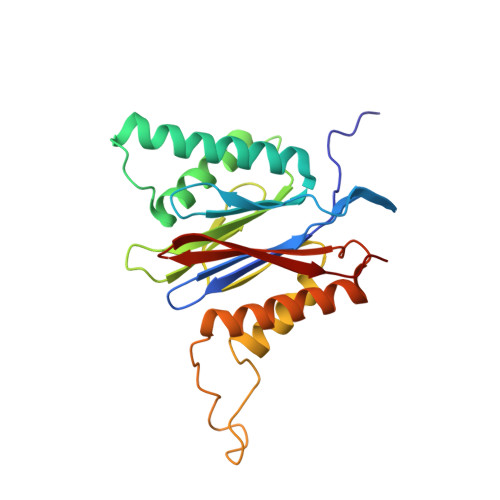
(-)-Homosalinosporamide A and Its Mode of Proteasome Inhibition: An X-ray Crystallographic Study.
Groll, M., Nguyen, H., Vellalath, S., Romo, D.(2018) Mar Drugs 16
- PubMed: 30029468
- DOI: https://doi.org/10.3390/md16070240
- Primary Citation of Related Structures:
6GOP - PubMed Abstract:
Upon acylation of the proteasome by the β-lactone inhibitor salinosporamide A (SalA), tetrahydrofuran formation occurs by intramolecular alkylation of the incipient alkoxide onto the choroethyl sidechain and irreversibly blocks the active site. Our previously described synthetic approach to SalA, utilizing a bioinspired, late-stage, aldol-β-lactonization strategy to construct the bicyclic β-lactone core, enabled synthesis of (⁻)-homosalinosporamide A (homoSalA). This homolog was targeted to determine whether an intramolecular tetrahydropyran is formed in a similar manner to SalA. Herein, we report the X-ray structure of the yeast 20S proteasome:homoSalA-complex which reveals that tetrahydropyran ring formation does not occur despite comparable potency at the chymotrypsin-like active site in a luminogenic enzyme assay. Thus, the natural product derivative homoSalA blocks the proteasome by a covalent reversible mode of action, opening the door for further fine-tuning of proteasome inhibition.
Organizational Affiliation:
Center for Integrated Protein Science Munich at the Department Chemie, Technische Universität München, Lichtenbergstrasse 4, 85747 Garching, Germany. michael.groll@tum.de.