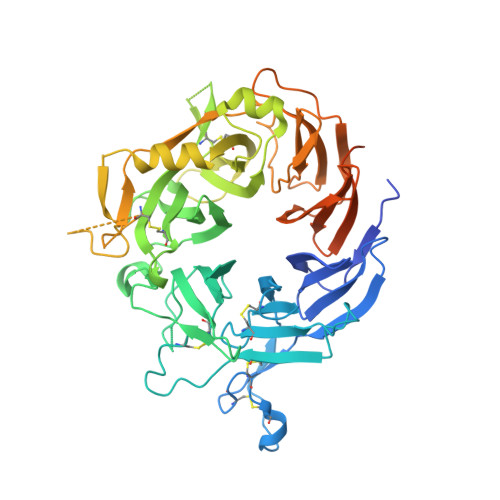
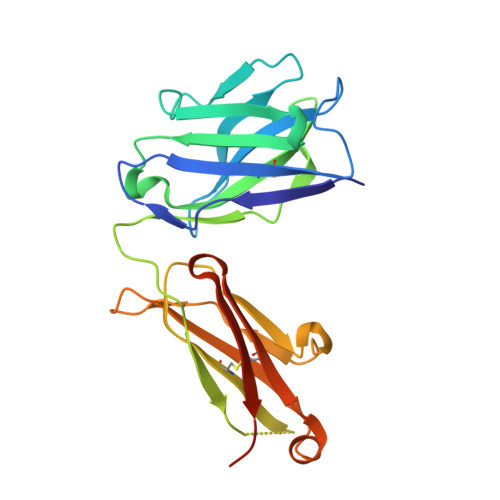
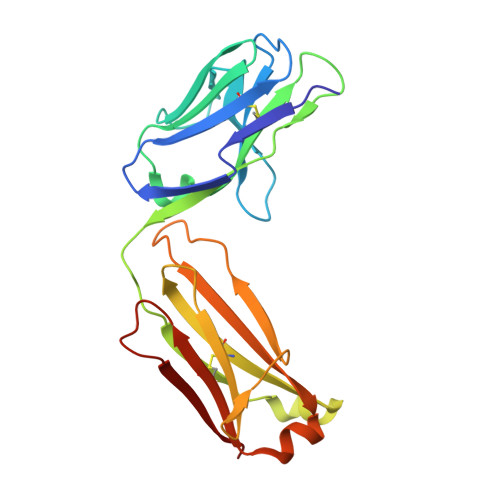
MM-131, a bispecific anti-Met/EpCAM mAb, inhibits HGF-dependent and HGF-independent Met signaling through concurrent binding to EpCAM.
Casaletto, J.B., Geddie, M.L., Abu-Yousif, A.O., Masson, K., Fulgham, A., Boudot, A., Maiwald, T., Kearns, J.D., Kohli, N., Su, S., Razlog, M., Raue, A., Kalra, A., Hakansson, M., Logan, D.T., Welin, M., Chattopadhyay, S., Harms, B.D., Nielsen, U.B., Schoeberl, B., Lugovskoy, A.A., MacBeath, G.(2019) Proc Natl Acad Sci U S A 116: 7533-7542
- PubMed: 30898885
- DOI: https://doi.org/10.1073/pnas.1819085116
- Primary Citation of Related Structures:
6HYG, 6I04, 6I07 - PubMed Abstract:
Activation of the Met receptor tyrosine kinase, either by its ligand, hepatocyte growth factor (HGF), or via ligand-independent mechanisms, such as MET amplification or receptor overexpression, has been implicated in driving tumor proliferation, metastasis, and resistance to therapy. Clinical development of Met-targeted antibodies has been challenging, however, as bivalent antibodies exhibit agonistic properties, whereas monovalent antibodies lack potency and the capacity to down-regulate Met. Through computational modeling, we found that the potency of a monovalent antibody targeting Met could be dramatically improved by introducing a second binding site that recognizes an unrelated, highly expressed antigen on the tumor cell surface. Guided by this prediction, we engineered MM-131, a bispecific antibody that is monovalent for both Met and epithelial cell adhesion molecule (EpCAM). MM-131 is a purely antagonistic antibody that blocks ligand-dependent and ligand-independent Met signaling by inhibiting HGF binding to Met and inducing receptor down-regulation. Together, these mechanisms lead to inhibition of proliferation in Met-driven cancer cells, inhibition of HGF-mediated cancer cell migration, and inhibition of tumor growth in HGF-dependent and -independent mouse xenograft models. Consistent with its design, MM-131 is more potent in EpCAM-high cells than in EpCAM-low cells, and its potency decreases when EpCAM levels are reduced by RNAi. Evaluation of Met, EpCAM, and HGF levels in human tumor samples reveals that EpCAM is expressed at high levels in a wide range of Met-positive tumor types, suggesting a broad opportunity for clinical development of MM-131.
Organizational Affiliation:
Discovery Division, Merrimack Pharmaceuticals, Inc., Cambridge, MA 02139.