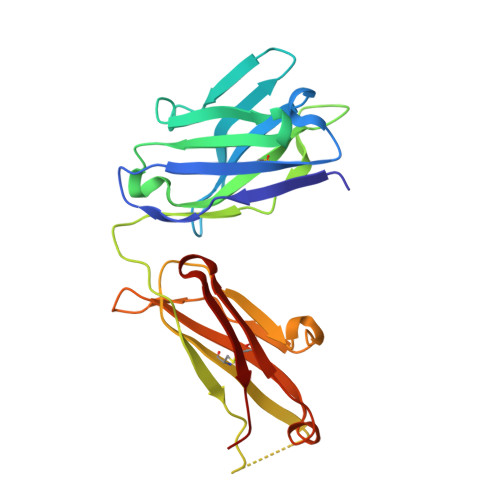
Structure-guided design fine-tunes pharmacokinetics, tolerability, and antitumor profile of multispecific frizzled antibodies.
Raman, S., Beilschmidt, M., To, M., Lin, K., Lui, F., Jmeian, Y., Ng, M., Fernandez, M., Fu, Y., Mascall, K., Duque, A., Wang, X., Pan, G., Angers, S., Moffat, J., Sidhu, S.S., Magram, J., Sinclair, A.M., Fransson, J., Julien, J.P.(2019) Proc Natl Acad Sci U S A 116: 6812-6817
- PubMed: 30894493
- DOI: https://doi.org/10.1073/pnas.1817246116
- Primary Citation of Related Structures:
6O39, 6O3A, 6O3B - PubMed Abstract:
Aberrant activation of Wnt/β-catenin signaling occurs frequently in cancer. However, therapeutic targeting of this pathway is complicated by the role of Wnt in stem cell maintenance and tissue homeostasis. Here, we evaluated antibodies blocking 6 of the 10 human Wnt/Frizzled (FZD) receptors as potential therapeutics. Crystal structures revealed a common binding site for these monoclonal antibodies (mAbs) on FZD, blocking the interaction with the Wnt palmitoleic acid moiety. However, these mAbs displayed gastrointestinal toxicity or poor plasma exposure in vivo. Structure-guided engineering was used to refine the binding of each mAb for FZD receptors, resulting in antibody variants with improved in vivo tolerability and developability. Importantly, the lead variant mAb significantly inhibited tumor growth in the HPAF-II pancreatic tumor xenograft model. Taken together, our data demonstrate that anti-FZD cancer therapeutic antibodies with broad specificity can be fine-tuned to navigate in vivo exposure and tolerability while driving therapeutic efficacy.
Organizational Affiliation:
Program in Molecular Medicine, Hospital for Sick Children Research Institute, Toronto, ON M5G 0A4, Canada.