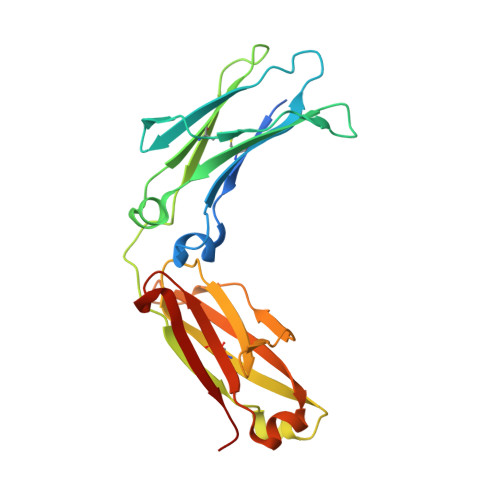
Structure and Dynamics of a Site-Specific Labeled Fc Fragment with Altered Effector Functions.
Gallagher, D.T., McCullough, C., Brinson, R.G., Ahn, J., Marino, J.P., Dimasi, N.(2019) Pharmaceutics 11
- PubMed: 31640157
- DOI: https://doi.org/10.3390/pharmaceutics11100546
- Primary Citation of Related Structures:
6P6D - PubMed Abstract:
Antibody-drug conjugates (ADCs) are a class of biotherapeutic drugs designed as targeted therapies for the treatment of cancer. Among the challenges in generating an effective ADC is the choice of an effective conjugation site on the IgG. One common method to prepare site-specific ADCs is to engineer solvent-accessible cysteine residues into antibodies. Here, we used X-ray diffraction and hydrogen-deuterium exchange mass spectroscopy to analyze the structure and dynamics of such a construct where a cysteine has been inserted after Ser 239 (Fc-239i) in the antibody heavy chain sequence. The crystal structure of this Fc-C239i variant at 0.23 nm resolution shows that the inserted cysteine structurally replaces Ser 239 and that this causes a domino-like backward shift of the local polypeptide, pushing Pro 238 out into the hinge. Proline is unable to substitute conformationally for the wild-type glycine at this position, providing a structural reason for the previously observed abolition of both FcγR binding and antibody-dependent cellular cytotoxicity. Energy estimates for the both the FcγR interface (7 kcal/mol) and for the differential conformation of proline (20 kcal/mol) are consistent with the observed disruption of FcγR binding, providing a quantifiable case where strain at a single residue appears to disrupt a key biological function. Conversely, the structure of Fc-C239i is relatively unchanged at the intersection of the CH2 and CH3 domains; the site known to be involved in binding of the neonatal Fc receptor (FcRn), and an alignment of the Fc-C239i structure with an Fc structure in a ternary Fc:FcRn:HSA (human serum albumin) complex implies that these favorable contacts would be maintained. Hydrogen deuterium exchange mass spectroscopy (HDX-MS) data further suggest a significant increase in conformational mobility for the Fc-C239i protein relative to Fc that is evident even far from the insertion site but still largely confined to the CH2 domain. Together, the findings provide a detailed structural and dynamic basis for previously observed changes in ADC functional binding to FcγR, which may guide further development of ADC designs.
Organizational Affiliation:
Institute for Bioscience and Biotechnology, National Institute of Standards and Technology and the University of Maryland, 9600 Gudelsky Drive, Rockville, MD 20850, USA. travis.gallagher@nist.gov.