Fast High-Pressure Freezing of Protein Crystals in Their Mother Liquor
Burkhardt, A., Warmer, M., Pannerselvam, S., Wagner, A., Zouni, A., Reimer, R., Hohenberg, H., Meents, A.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 495
- PubMed: 22505429
- DOI: https://doi.org/10.1107/S1744309112009670
- Primary Citation of Related Structures:
4A7D, 4A7E - PubMed Abstract:
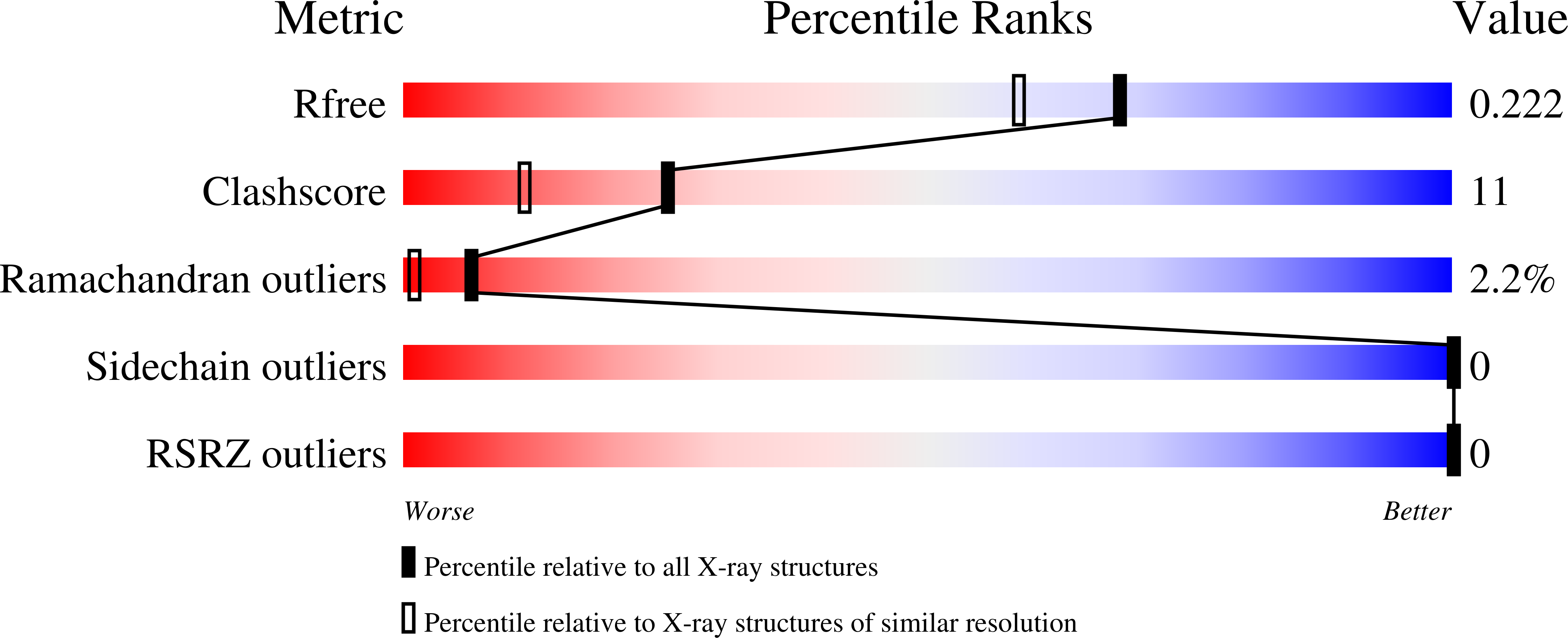
High-pressure freezing (HPF) is a method which allows sample vitrification without cryoprotectants. In the present work, protein crystals were cooled to cryogenic temperatures at a pressure of 210 MPa. In contrast to other HPF methods published to date in the field of cryocrystallography, this protocol involves rapid sample cooling using a standard HPF device. The fast cooling rates allow HPF of protein crystals directly in their mother liquor without the need for cryoprotectants or external reagents. HPF was first attempted with hen egg-white lysozyme and cubic insulin crystals, yielding good to excellent diffraction quality. Non-cryoprotected crystals of the membrane protein photosystem II have been successfully cryocooled for the first time. This indicates that the presented HPF method is well suited to the vitrification of challenging systems with large unit cells and weak crystal contacts.
Organizational Affiliation:
Deutsches Elektronen-Synchrotron DESY, Notkestrasse 85, 22607 Hamburg, Germany. anja.burkhardt@desy.de