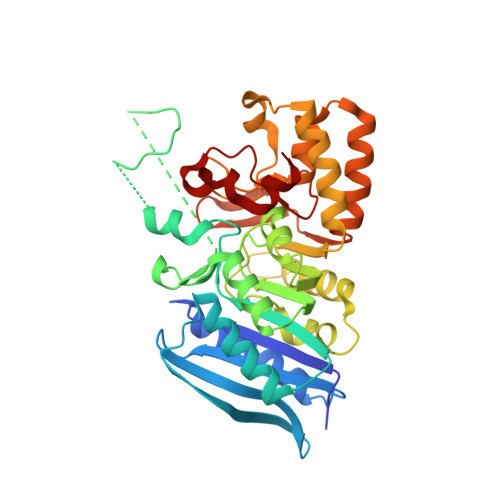
X-ray Structures of Two Bacteroides thetaiotaomicron C11 Proteases in Complex with Peptide-Based Inhibitors.
Roncase, E.J., Gonzalez-Paez, G.E., Wolan, D.W.(2019) Biochemistry 58: 1728-1737
- PubMed: 30835452
- DOI: https://doi.org/10.1021/acs.biochem.9b00098
- Primary Citation of Related Structures:
6N9J, 6NAG - PubMed Abstract:
Commensal bacteria secrete proteins and metabolites to influence host intestinal homeostasis, and proteases represent a significant constituent of the components at the host:microbiome interface. Here, we determined the structures of the two secreted C11 cysteine proteases encoded by the established gut commensal Bacteroides thetaiotaomicron. We employed mutational analysis to demonstrate the two proteases, termed "thetapain" and "iotapain", undergo in trans autoactivation after lysine and/or arginine residues, as observed for other C11 proteases. We determined the structures of the active forms of thetapain and iotapain in complex with irreversible peptide inhibitors, Ac-VLTK-AOMK and biotin-VLTK-AOMK, respectively. Structural comparisons revealed key active-site interactions important for peptide recognition are more extensive for thetapain; however, both proteases employ a glutamate residue to preferentially bind small polar residues at the P2 position. Our results will aid in the design of protease-specific probes to ultimately understand the biological role of C11 proteases in bacterial fitness, elucidate their host and/or microbial substrates, and interrogate their involvement in microbiome-related diseases.
Organizational Affiliation:
Departments of Molecular Medicine and Integrative Structural and Computational Biology , The Scripps Research Institute , 10550 North Torrey Pines Road , La Jolla , California 92037 , United States.