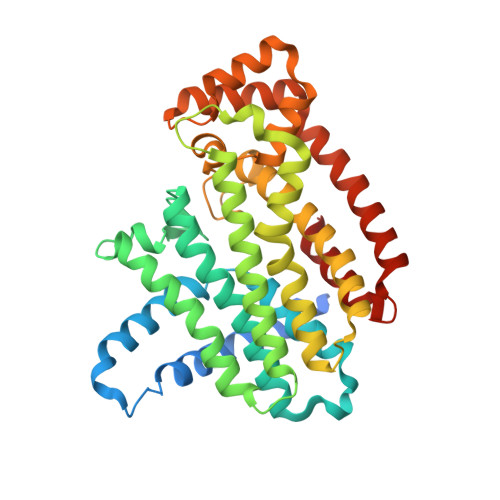
Chirality-Driven Mode of Binding of alpha-Aminophosphonic Acid-Based Allosteric Inhibitors of the Human Farnesyl Pyrophosphate Synthase (hFPPS).
Feng, Y., Park, J., Li, S.G., Boutin, R., Viereck, P., Schilling, M.A., Berghuis, A.M., Tsantrizos, Y.S.(2019) J Med Chem 62: 9691-9702
- PubMed: 31577901
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01104
- Primary Citation of Related Structures:
6N7Y, 6N7Z, 6N82, 6N83, 6OAG, 6OAH - PubMed Abstract:
Thienopyrimidine-based allosteric inhibitors of the human farnesyl pyrophosphate synthase (hFPPS), characterized by a chiral α-aminophosphonic acid moiety, were synthesized as enantiomerically enriched pairs, and their binding mode was investigated by X-ray crystallography. A general consensus in the binding orientation of all ( R )- and ( S )-enantiomers was revealed. This finding is a prerequisite for establishing a reliable structure-activity relationship (SAR) model.
Organizational Affiliation:
Department of Chemistry , McGill University , 801 Sherbrooke Street West , Montreal , Quebec H3A 0B8 , Canada.