Full Text |
QUERY: Gene Name = "Ta1" | MyPDB Login | Search API |
| Search Summary | This query matches 20 Structures. |
Structure Determination MethodologyScientific Name of Source OrganismTaxonomyExperimental MethodPolymer Entity TypeRefinement Resolution (Å)Release DateEnzyme Classification NameMembrane Protein AnnotationSymmetry Type | 1 to 20 of 20 Structures Page 1 of 1 Sort by
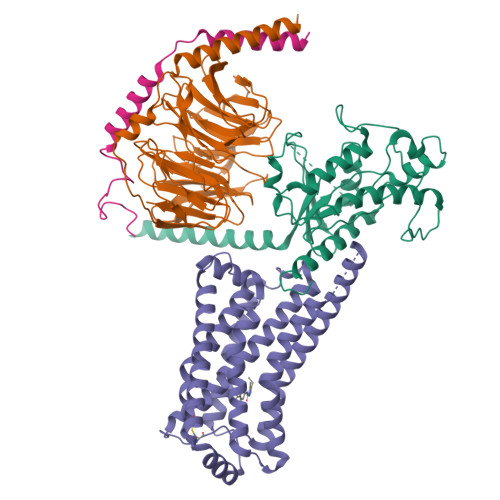
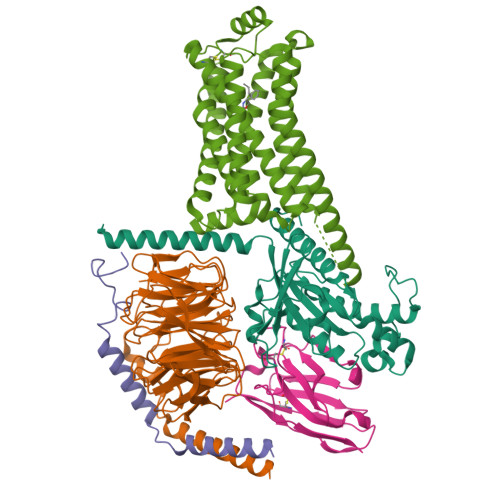
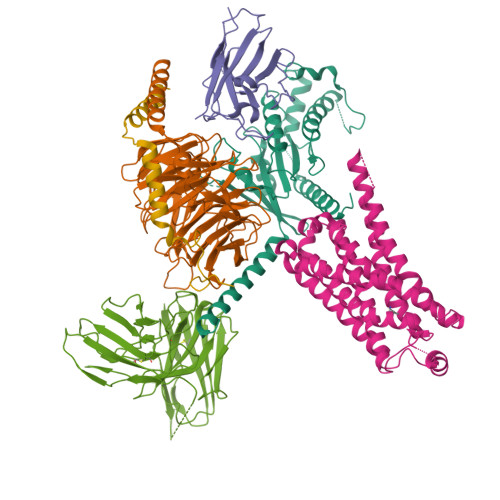
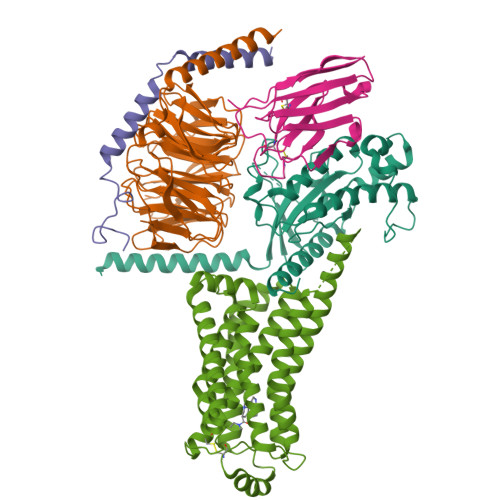
Ralmitaront(RO-6889450)-bound hTAAR1-Gs protein complexXu, Z., Guo, L.L., Zhao, C., Shen, S.Y., Sun, J.P., Shao, Z.H. (2023) Nature 624: 672-681
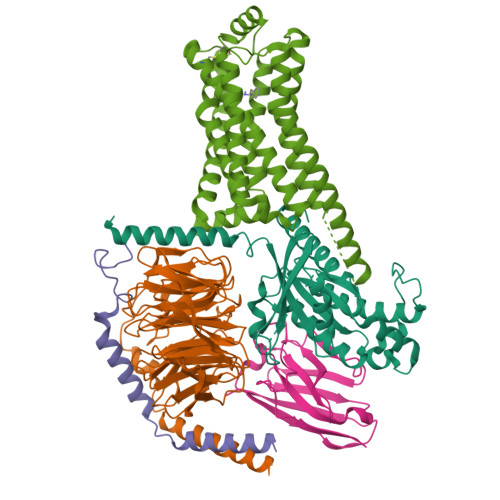
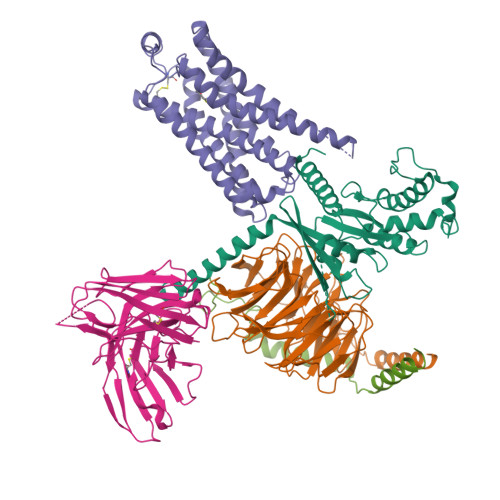
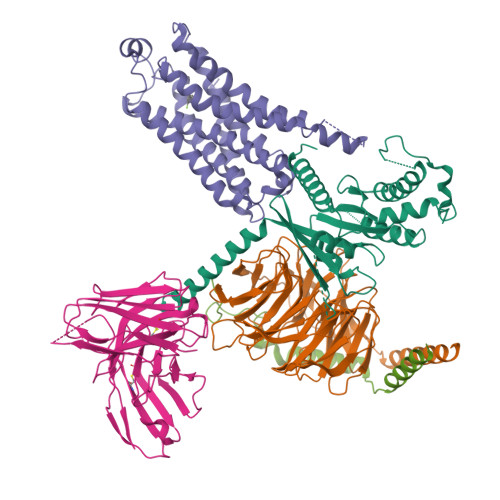
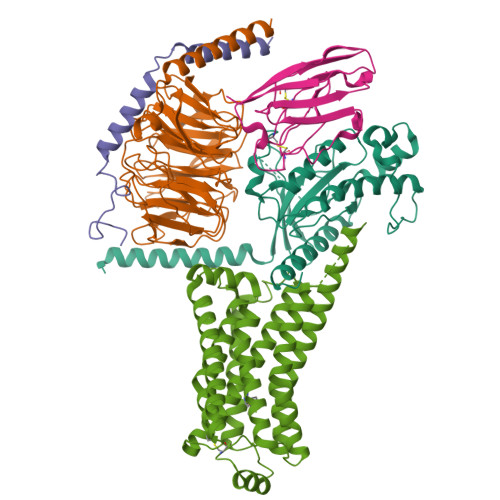
Cryo-EM structure of the METH-TAAR1 complexLiu, H., Zheng, Y., Wang, Y., Wang, Y., He, X., Xu, P., Huang, S., Yuan, Q., Zhang, X., Wang, S., Xu, H.E., Xu, F. (2023) Nature 624: 663-671
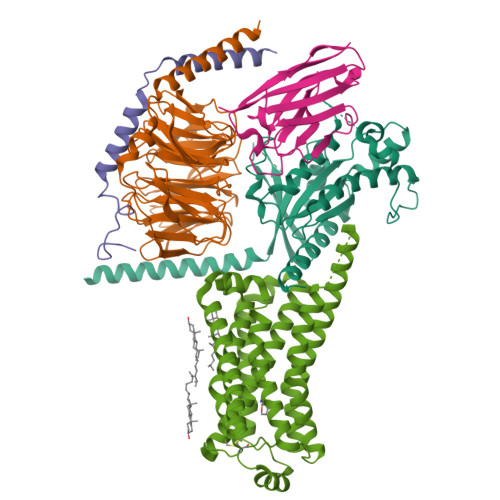
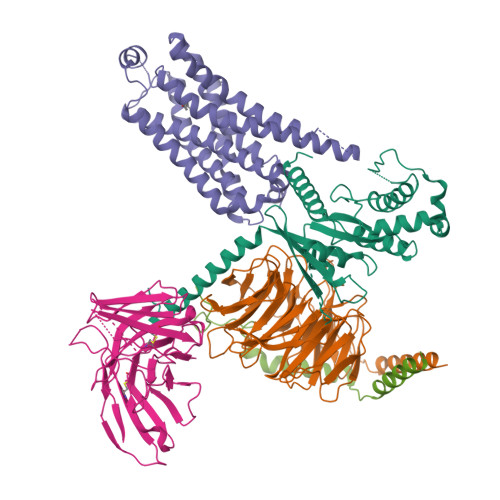
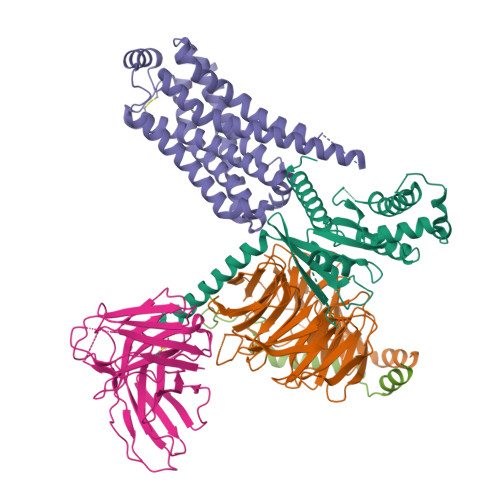
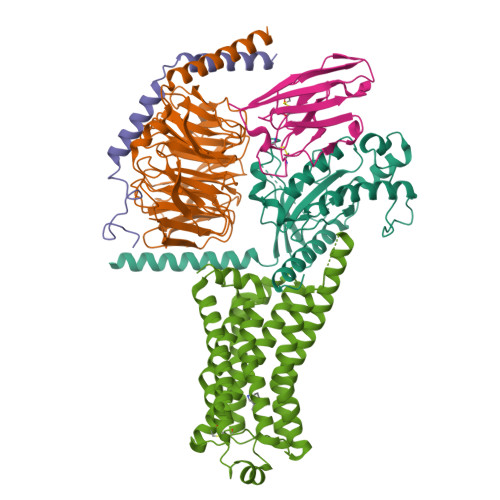
Cryo-EM structure of the SEP363856-bound TAAR1-Gs complexLiu, H., Zheng, Y., Wang, Y., Wang, Y., He, X., Xu, P., Huang, S., Yuan, Q., Zhang, X., Wang, S., Xu, H.E., Xu, F. (2023) Nature 624: 663-671
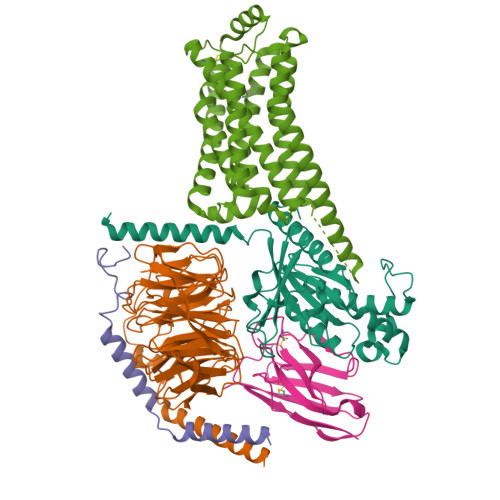
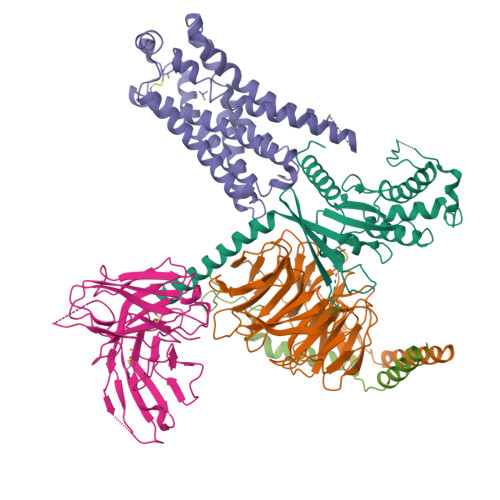
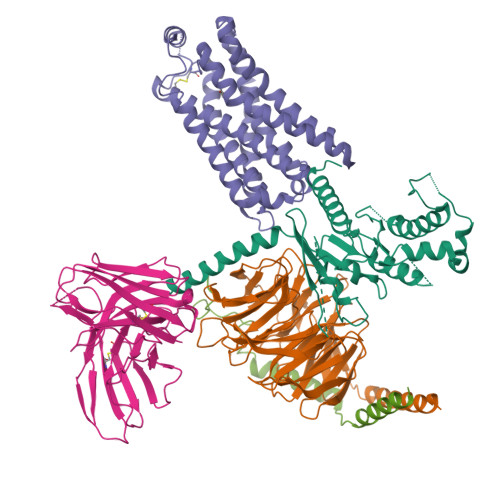
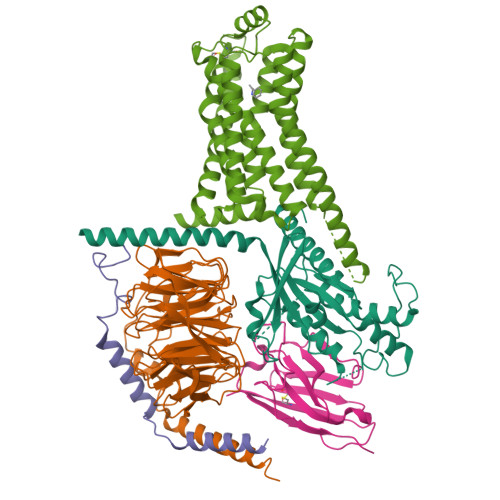
Cryo-EM structure of the PEA-bound TAAR1-Gs complexLiu, H., Zheng, Y., Wang, Y., Wang, Y., He, X., Xu, P., Huang, S., Yuan, Q., Zhang, X., Wang, S., Xu, H.E., Xu, F. (2023) Nature 624: 663-671
Cryo-EM structure of the RO5256390-TAAR1 complexLiu, H., Zheng, Y., Wang, Y., Wang, Y., He, X., Xu, P., Huang, S., Yuan, Q., Zhang, X., Wang, S., Xu, H.E., Xu, F. (2023) Nature 624: 663-671
Cryo-EM structure of the SEP363856-bound mTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the ZH8651-bound mTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the TMA-bound mTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the PEA-bound mTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the ZH8667-bound mTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the ZH8651-bound hTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the ZH8651-bound mTAAR1-Gq complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the PEA-bound hTAAR1-Gs complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the CHA-bound mTAAR1-Gq complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the CHA-bound mTAAR1 complexRong, N.K., Guo, L.L., Zhang, M.H., Li, Q., Yang, F., Sun, J.P. (2023) Cell 186: 5347-5362.e24
Cryo-EM structure of the apo hTAAR1-Gs complexJiang, K.X., Zheng, Y., Xu, F. (2024) Cell Rep 43: 114505-114505
Cryo-EM structure of the LSD-bound hTAAR1-Gs complexJiang, K.X., Zheng, Y., Xu, F. (2024) Cell Rep 43: 114505-114505
Cryo-EM structure of the RO5263397-bound hTAAR1-Gs complexJiang, K.X., Zheng, Y., Xu, F. (2024) Cell Rep 43: 114505-114505
Cryo-EM structure of the RO5263397-bound mTAAR1-Gs complexJiang, K.X., Zheng, Y., Xu, F. (2024) Cell Rep 43: 114505-114505
Cryo-EM structure of the METH-bound hTAAR1-Gs complex(2024) J Med Chem 67: 18593-18605
1 to 20 of 20 Structures Page 1 of 1 Sort by |