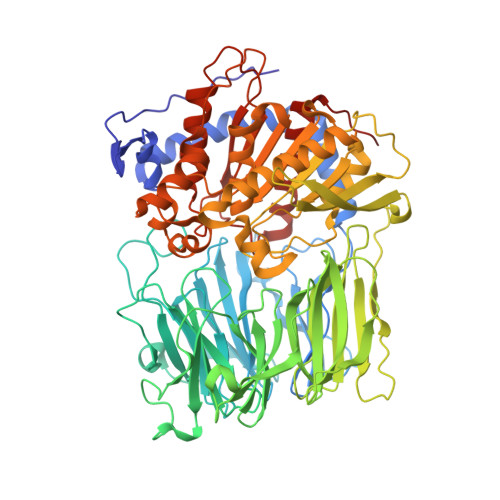
Structures of Prolyl Oligopeptidase Substrate/ Inhibitor Complexes. Use of Inhibitor Binding for Titration of the Catalytic Histidine Residue
Fulop, V., Szeltner, Z., Renner, V., Polgar, L.(2001) J Biological Chem 276: 1262
- PubMed: 11031266 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M007003200
- Primary Citation Related Structures:
1E8M, 1E8N - PubMed Abstract:
Structure determination of the inactive S554A variant of prolyl oligopeptidase complexed with an octapeptide has shown that substrate binding is restricted to the P4-P2' region. In addition, it has revealed a hydrogen bond network of potential catalytic importance not detected in other serine peptidases. This involves a unique intramolecular hydrogen bond between the P1' amide and P2 carbonyl groups and another between the P2' amide and Nepsilon2 of the catalytic histidine 680 residue. It is argued that both hydrogen bonds promote proton transfer from the imidazolium ion to the leaving group. Another complex formed with the product-like inhibitor benzyloxycarbonyl-glycyl-proline, indicating that the carboxyl group of the inhibitor forms a hydrogen bond with the Nepsilon2 of His(680). Because a protonated histidine makes a stronger interaction with the carboxyl group, it offers a possibility of the determination of the real pK(a) of the catalytic histidine residue. This was found to be 6.25, lower than that of the well studied serine proteases. The new titration method gave a single pK(a) for prolyl oligopeptidase, whose reaction exhibited a complex pH dependence for k(cat)/K(m), and indicated that the observed pK(a) values are apparent. The procedure presented may be applicable for other serine peptidases.
- Department of Biological Sciences, University of Warwick, Gibbet Hill Road, Coventry CV4 7AL, United Kingdom.
Organizational Affiliation: