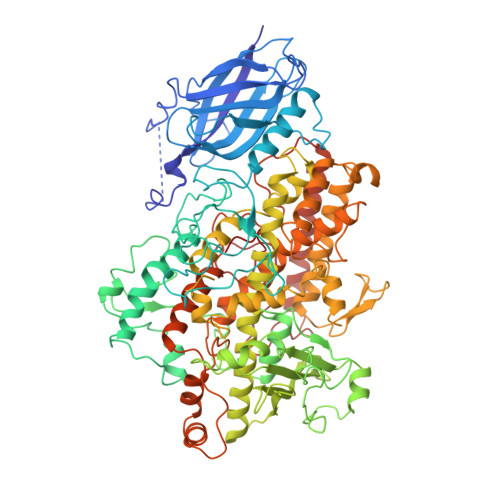
Structure of soybean lipoxygenase L3 and a comparison with its L1 isoenzyme.
Skrzypczak-Jankun, E., Amzel, L.M., Kroa, B.A., Funk Jr., M.O.(1997) Proteins 29: 15-31
- PubMed: 9294864 Search on PubMed
- Primary Citation Related Structures:
1LNH - PubMed Abstract:
Soybean lipoxygenase isoenzyme L3 represents a second example (after L1) of the X-ray structure (R = 17% at 2.6 A resolution) for a member of the large family of lipoxygenases. L1 and L3 have different characteristics in catalysis, although they share 72% sequence identity (the changes impact 255 amino acids) and similar folding (average C alpha rms deviation of 1 A). The critical nonheme iron site has the same features as for L1:3O and 3N in pseudo C3v orientation, with two oxygen atoms (from Asn713 and water) at a nonbinding distance. Asn713 and His518 are strategically located at the junction of three cavities connecting the iron site with the molecule surface. The most visible differences between L1 and L3 isoenzymes occur in and near these cavities, affecting their accessibility and volume. Among the L1/L3 substitutions Glu256/ Thr274, Tyr409/His429, and Ser747/Asp766 affect the salt bridges (L1: Glu256...His248 and Asp490...Arg707) that in L1 restrict the access to the iron site from two opposite directions. The L3 molecule has a passage going through the whole length of the helical domain, starting at the interface with the Nt-domain (near 25-27 and 254-278) and going to the opposite end of the Ct-domain (near 367, 749). The substrate binding and the role of His513, His266, His776 (and other residues nearby) are illustrated and discussed by using models of linoleic acid binding. These hypotheses provide a possible explanation for a stringent stereo-specificity of catalytic products in L1 (that produces predominantly 13-hydroperoxide) versus the lack of such specificity in L3 (that turns out a mixture of 9- and 13-hydroperoxides and their diastereoisomers).
- Department of Chemistry, University of Toledo, Ohio 43606, USA. ejankun@uoft02.utoledo.edu
Organizational Affiliation: