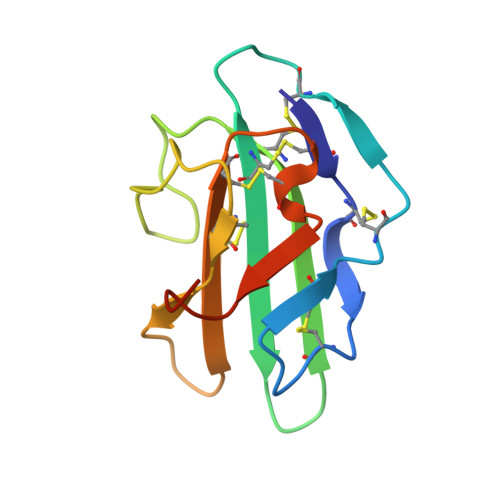
THE 1.1A CRYSTAL STRUCTURE OF HUMAN TGF-BETA TYPE II RECEPTOR LIGAND BINDING DOMAIN
Boesen, C.C., Radaev, S., Motyka, S.A., Patamawenu, A., Sun, P.D.(2002) Structure 10: 913-919
- PubMed: 12121646 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00780-3
- Primary Citation Related Structures:
1M9Z - PubMed Abstract:
Transforming growth factor beta (TGF-beta) is involved in a wide range of biological functions including development, carcinogenesis, and immune regulation. Here we report the 1.1 A resolution crystal structure of human TGF-beta type II receptor ectodomain (TBRII). The overall structure of TBRII is similar to that of activin type II receptor ectodomain (ActRII) and bone morphogenic protein receptor type IA (BRIA). It displays a three-finger toxin fold with fingers formed by the beta strand pairs beta1-beta2, beta3-beta4, and beta5-beta6. The first finger in the TBRII is significantly longer than in ActRII and BRIA and folds tightly between the second finger and the C terminus. Surface charge distributions and hydrophobic patches predict potential TBRII binding sites.
- Structural Biology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, MD 20852, USA.
Organizational Affiliation: