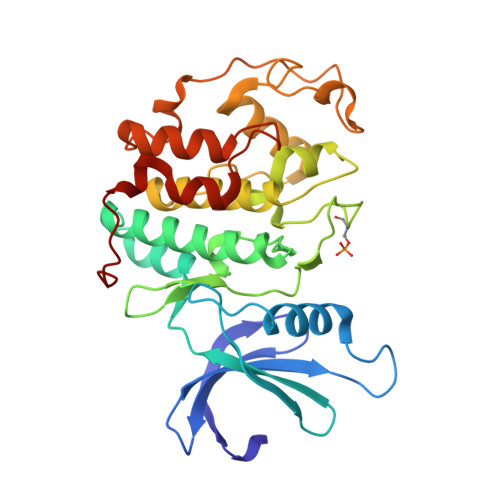
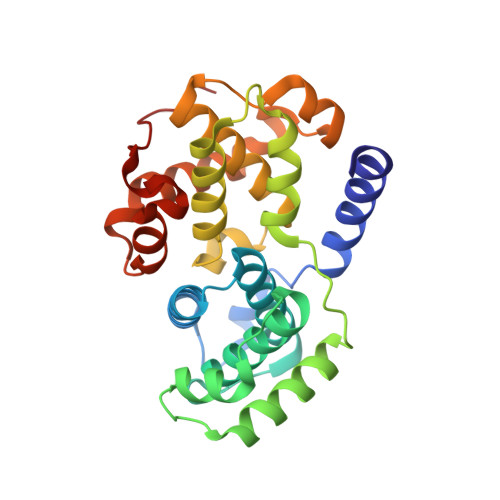
Alternative binding modes of an inhibitor to two different kinases
De Moliner, E., Brown, N.R., Johnson, L.N.(2003) Eur J Biochem 270: 1-8
- PubMed: 12869192
- DOI: https://doi.org/10.1046/j.1432-1033.2003.03697.x
- Primary Citation of Related Structures:
1P5E - PubMed Abstract:
Protein kinases are targets for therapeutic agents designed to intervene in signaling processes in the diseased state. Most kinase inhibitors are directed towards the conserved ATP binding site. Because the essential features of this site are conserved in all eukaryotic protein kinases, it is generally assumed that the same compound will bind in a similar manner to different protein kinases. The inhibitor 4,5,6,7-tetrabromobenzotriazole (TBB) is a selective inhibitor for the protein kinase CK2 (IC50 1.6 micro m) (Sarno et al. (2001) FEBS Letts.496, 44-48). Three other kinases [cyclin-dependent protein kinase 2 (CDK2), phosphorylase kinase and glycogen synthase kinase 3beta] exhibit approximately 10-fold weaker affinity for TBB than CK2. We report the crystal structure of TBB in complex with phospho-CDK2-cyclin A at 2.2 A resolution and compare the interactions with those observed for TBB bound to CK2. TBB binds at the ATP binding site of both kinases. In CDK2, each of the four bromine atoms makes polar contacts either to main chain oxygens in the hinge region of the kinase or to water molecules, in addition to several van der Waals contacts. The mode of binding of TBB to CDK2 is different from that to CK2. TBB in CDK2 is displaced more towards the hinge region between the N- and C-terminal lobes and rotated relative to TBB in CK2. The ATP binding pocket is wider in CDK2 than in CK2 resulting in fewer van der Waals contacts but TBB in CK2 does not contact the hinge. The structures show that, despite the conservation of the ATP binding pocket, the inhibitor is able to exploit different recognition features so that the same compound can bind in different ways to the two different kinases.
- Laboratory of Molecular Biophysics, Department of Biochemistry, University of Oxford, UK.
Organizational Affiliation: