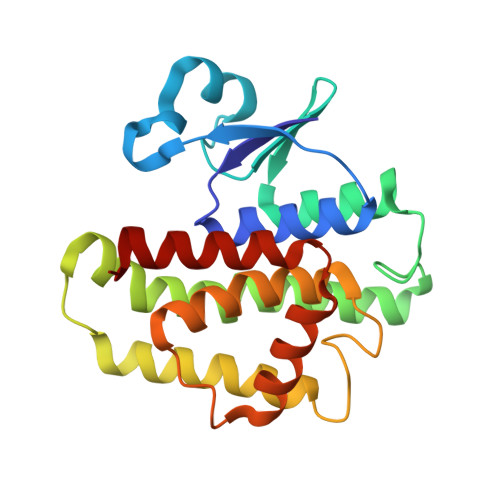
Structure of an insect delta-class glutathione S-transferase from a DDT-resistant strain of the malaria vector Anopheles gambiae.
Chen, L., Hall, P.R., Zhou, X.E., Ranson, H., Hemingway, J., Meehan, E.J.(2003) Acta Crystallogr D Biol Crystallogr 59: 2211-2217
- PubMed: 14646079 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903018493
- Primary Citation Related Structures:
1PN9 - PubMed Abstract:
Glutathione S-transferases (GSTs) are a major family of detoxification enzymes which possess a wide range of substrate specificities. Most organisms possess many GSTs belonging to multiple classes. Interest in GSTs in insects is focused on their role in insecticide resistance; many resistant insects have elevated levels of GST activity. In the malaria vector Anopheles gambiae, elevated GST levels are associated with resistance to the organochlorine insecticide DDT [1,1,1-trichloro-2,2-bis-(p-chlorophenyl)ethane]. This mosquito is the source of an insect GST, agGSTd1-6, which metabolizes DDT and is inhibited by a number of pyrethroid insecticides. The crystal structure of agGSTd1-6 in complex with its inhibitor S-hexyl glutathione has been determined and refined at 2.0 A resolution. The structure adopts a classical GST fold and is similar to those of other insect delta-class GSTs, implying a common conjugation mechanism. A structure-based model for the binding of DDT to agGSTd1-6 reveals two subpockets in the hydrophobic binding site (H-site), each accommodating one planar p-chlorophenyl ring.
- Laboratory for Structural Biology, Department of Chemistry, Graduate Programs of Biotechnology, Chemistry and Materials Science, University of Alabama in Huntsville, Huntsville, AL 35899, USA. chenlq@email.uah.edu
Organizational Affiliation: