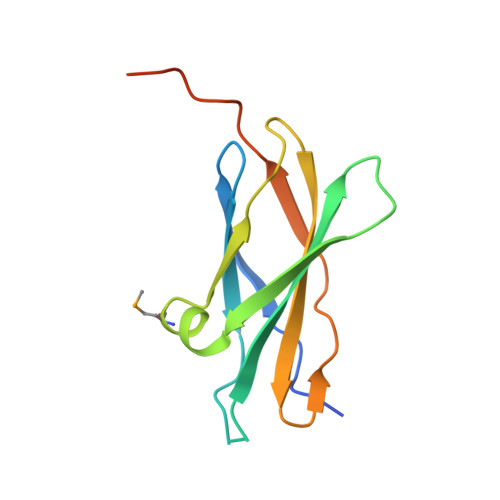
Crystal structure of the first fibronectin type III domain of neural cell adhesion molecule splicing isoform from human muscle culture lambda-4.4
Saijo, S., Nishino, A., Kishishita, S., Chen, L., Liu, Z.J., Wang, B.C., Shirouzu, M., Yokoyama, S.To be published.