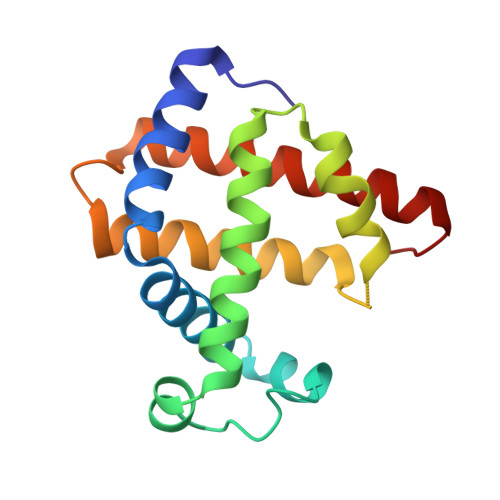
Design and Structure Analysis of Artificial Metalloproteins: Selective Coordination of His64 to Copper Complexes with Square-Planar Structure in the apo-Myoglobin Scaffold
Abe, S., Ueno, T., Reddy, P.A.N., Okazaki, S., Hikage, T., Suzuki, A., Yamane, T., Nakajima, H., Watanabe, Y.(2007) Inorg Chem 46: 5137-5139
- PubMed: 17523632 Search on PubMed
- DOI: https://doi.org/10.1021/ic070289m
- Primary Citation Related Structures:
2EB8, 2EB9 - PubMed Abstract:
apo-Myoglobin (apo-Mb) was reconstituted with three copper complexes: CuII(Sal-Phe) (1; Sal-Phe = N-salicylidene-L-phenylalanato), CuII(Sal-Leu) (2; Sal-Leu = N-salicylidene-L-leucinato), and CuII(Sal-Ala) (3; Sal-Ala = N-salicylidene-L-alanato). The crystal structures of 1.apo-Mb (1.65 Angstrom resolution) and 2.apo-Mb (1.8 Angstrom resolution) show that the coordination geometry around the CuII atom in apo-Mb is distorted square-planar with tridentate Sal-X and a Nepsilon atom of His64 in the apo-Mb cavity and the plane of these copper complexes is perpendicular to that of heme. These results suggest that the apo-Mb cavity can hold metal complexes with various coordination geometries.
- Department of Chemistry, Graduate School of Science, Nagoya University, Nagoya 464-8602, Japan.
Organizational Affiliation: