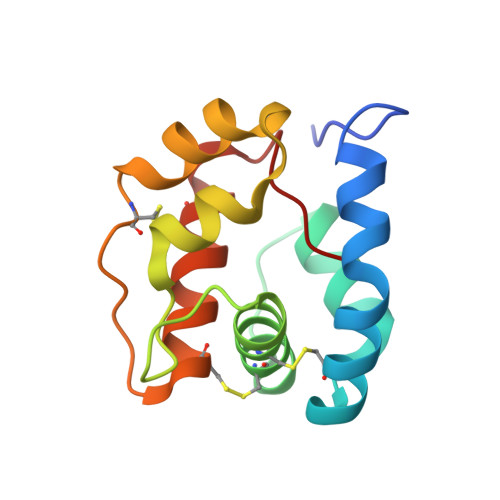
The crystal structure of an odorant binding protein from Anopheles gambiae: Evidence for a common ligand release mechanism.
Wogulis, M., Morgan, T., Ishida, Y., Leal, W.S., Wilson, D.K.(2006) Biochem Biophys Res Commun 339: 157-164
- PubMed: 16300742 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2005.10.191
- Primary Citation Related Structures:
2ERB - PubMed Abstract:
The Anopheles gambiae mosquito is the main vector of malaria transmission in sub-Saharan Africa. We present here a 1.5A crystal structure of AgamOBP1, an odorant binding protein (OBP) from the A. gambiae mosquito. The protein crystallized as a dimer with a unique binding pocket consisting of a continuous tunnel running through both subunits of the dimer and occupied by a PEG molecule. We demonstrate that AgamOBP1 undergoes a pH dependent conformational change that is associated with reduced ligand binding. A predominance of acid-labile hydrogen bonds involving the C-terminal loop suggests a mechanism in which a drop in pH causes C-terminal loop to open, leaving the binding tunnel solvent exposed, thereby lowering binding affinity for ligand. Because proteins from two distantly related insects also undergo a pH dependent conformational change involving the C-terminus that is associated with reduced ligand affinity, our results suggest a common mechanism for OBP activity.
- Section of Molecular and Cellular Biology, University of California, Davis, CA 95616, USA.
Organizational Affiliation: