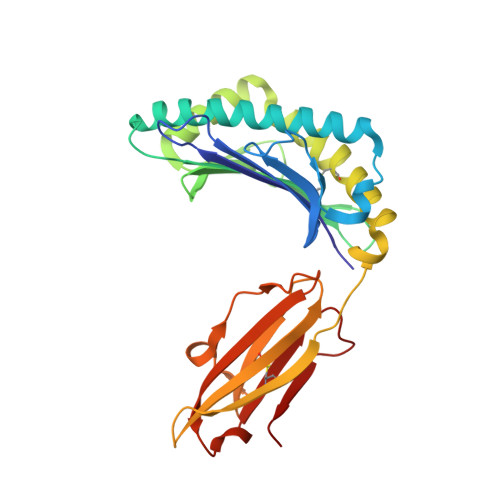
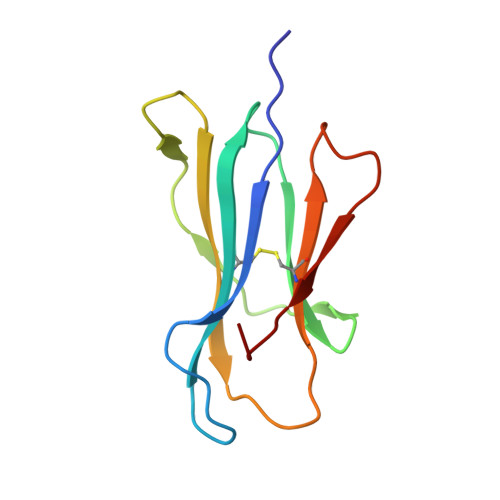
Structures of MART-1(26/27-35) Peptide/HLA-A2 Complexes Reveal a Remarkable Disconnect between Antigen Structural Homology and T Cell Recognition
Borbulevych, O.Y., Insaidoo, F.K., Baxter, T.K., Powell, D.J., Johnson, L.A., Restifo, N.P., Baker, B.M.(2007) J Mol Biology 372: 1123-1136
- PubMed: 17719062 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.07.025
- Primary Citation Related Structures:
2GT9, 2GTW, 2GTZ, 2GUO - PubMed Abstract:
Small structural changes in peptides presented by major histocompatibility complex (MHC) molecules often result in large changes in immunogenicity, supporting the notion that T cell receptors are exquisitely sensitive to antigen structure. Yet there are striking examples of TCR recognition of structurally dissimilar ligands. The resulting unpredictability of how T cells will respond to different or modified antigens impacts both our understanding of the physical bases for TCR specificity as well as efforts to engineer peptides for immunomodulation. In cancer immunotherapy, epitopes and variants derived from the MART-1/Melan-A protein are widely used as clinical vaccines. Two overlapping epitopes spanning amino acid residues 26 through 35 are of particular interest: numerous clinical studies have been performed using variants of the MART-1 26-35 decamer, although only the 27-35 nonamer has been found on the surface of targeted melanoma cells. Here, we show that the 26-35 and 27-35 peptides adopt strikingly different conformations when bound to HLA-A2. Nevertheless, clonally distinct MART-1(26/27-35)-reactive T cells show broad cross-reactivity towards these ligands. Simultaneously, however, many of the cross-reactive T cells remain unable to recognize anchor-modified variants with very subtle structural differences. These dichotomous observations challenge our thinking about how structural information on unligated peptide/MHC complexes should be best used when addressing questions of TCR specificity. Our findings also indicate that caution is warranted in the design of immunotherapeutics based on the MART-1 26/27-35 epitopes, as neither cross-reactivity nor selectivity is predictable based on the analysis of the structures alone.
- Department of Chemistry and Biochemistry, 251 Nieuwland Science Hall, University of Notre Dame, Notre Dame, IN 46556, USA.
Organizational Affiliation: