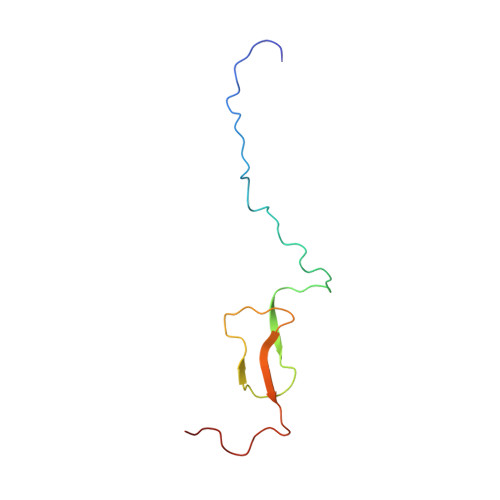
A surface loop directs conformational switching of a lipoyl domain between a folded and a novel misfolded structure.
Stott, K.M., Yusof, A.M., Perham, R.N., Jones, D.D.(2009) Structure 17: 1117-1127
- PubMed: 19679089 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2009.07.001
- Primary Citation Related Structures:
2K7V - PubMed Abstract:
A prominent surface loop links the first two beta strands of the lipoyl domain (E2plip) from the pyruvate dehydrogenase multienzyme complex of Escherichia coli. We show here that shortening this loop by two residues generates a protein that populates two structurally distinct stable conformers: an active, native-like monomer (HM) and a functionally compromised misfolded dimer (LM). Conversion of LM to HM was observed after exposure to temperatures above 50 degrees C. Removal of two additional residues from the loop caused the protein to adopt exclusively the misfolded conformation. Detailed NMR structural studies of the misfolded dimer reveal that the N-terminal half of the domain was unfolded and dynamic, whereas the C-terminal halves of two monomers had associated to form a structure with two-fold symmetry and a topology mimicking that of the folded monomer. The surface loop is therefore a hitherto unsuspected determinant in the folding process that leads to a functional protein.
- Department of Biochemistry, University of Cambridge, Cambridge, UK.
Organizational Affiliation: