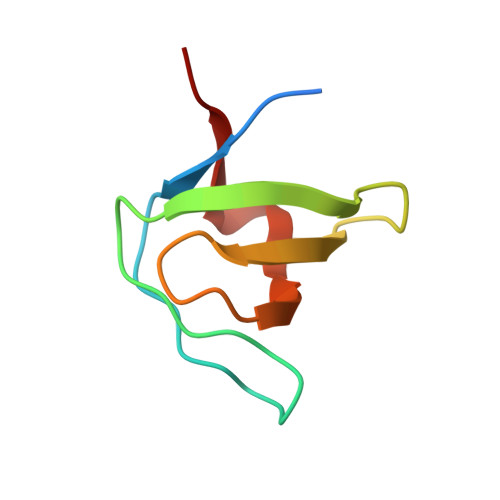
Structure of an intermediate state in protein folding and aggregation.
Neudecker, P., Robustelli, P., Cavalli, A., Walsh, P., Lundstrom, P., Zarrine-Afsar, A., Sharpe, S., Vendruscolo, M., Kay, L.E.(2012) Science 336: 362-366
- PubMed: 22517863 Search on PubMed
- DOI: https://doi.org/10.1126/science.1214203
- Primary Citation Related Structures:
2L2P, 2LP5 - PubMed Abstract:
Protein-folding intermediates have been implicated in amyloid fibril formation involved in neurodegenerative disorders. However, the structural mechanisms by which intermediates initiate fibrillar aggregation have remained largely elusive. To gain insight, we used relaxation dispersion nuclear magnetic resonance spectroscopy to determine the structure of a low-populated, on-pathway folding intermediate of the A39V/N53P/V55L (A, Ala; V, Val; N, Asn; P, Pro; L, Leu) Fyn SH3 domain. The carboxyl terminus remains disordered in this intermediate, thereby exposing the aggregation-prone amino-terminal β strand. Accordingly, mutants lacking the carboxyl terminus and thus mimicking the intermediate fail to safeguard the folding route and spontaneously form fibrillar aggregates. The structure provides a detailed characterization of the non-native interactions stabilizing an aggregation-prone intermediate under native conditions and insight into how such an intermediate can derail folding and initiate fibrillation.
- Department of Biochemistry, University of Toronto, Toronto, Ontario M5S 1A8, Canada.
Organizational Affiliation: