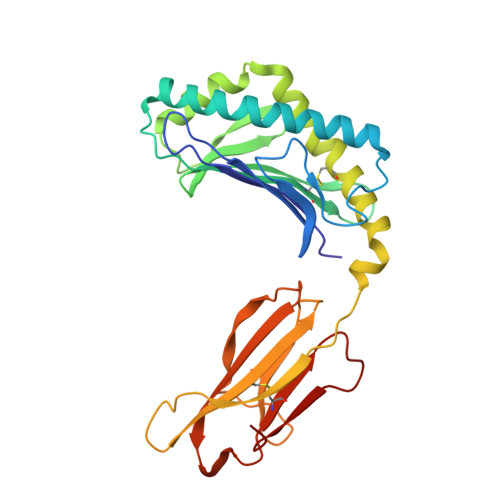
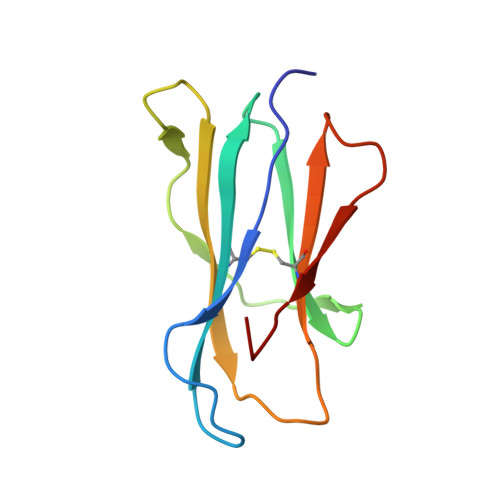
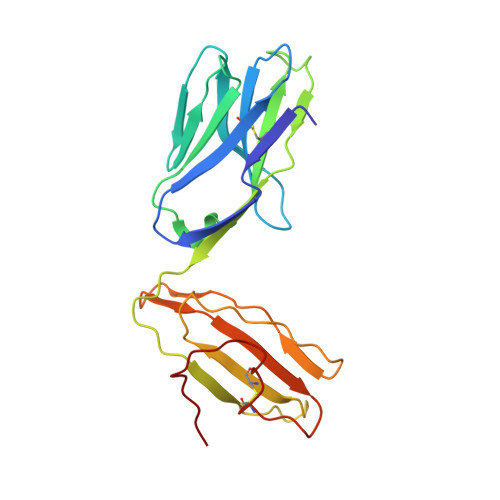
CD1d-lipid-antigen recognition by the semi-invariant NKT T-cell receptor.
Borg, N.A., Wun, K.S., Kjer-Nielsen, L., Wilce, M.C., Pellicci, D.G., Koh, R., Besra, G.S., Bharadwaj, M., Godfrey, D.I., McCluskey, J., Rossjohn, J.(2007) Nature 448: 44-49
- PubMed: 17581592
- DOI: https://doi.org/10.1038/nature05907
- Primary Citation of Related Structures:
2PO6 - PubMed Abstract:
The CD1 family is a large cluster of non-polymorphic, major histocompatibility complex (MHC) class-I-like molecules that bind distinct lipid-based antigens that are recognized by T cells. The most studied group of T cells that interact with lipid antigens are natural killer T (NKT) cells, which characteristically express a semi-invariant T-cell receptor (NKT TCR) that specifically recognizes the CD1 family member, CD1d. NKT-cell-mediated recognition of the CD1d-antigen complex has been implicated in microbial immunity, tumour immunity, autoimmunity and allergy. Here we describe the structure of a human NKT TCR in complex with CD1d bound to the potent NKT-cell agonist alpha-galactosylceramide, the archetypal CD1d-restricted glycolipid. In contrast to T-cell receptor-peptide-antigen-MHC complexes, the NKT TCR docked parallel to, and at the extreme end of the CD1d-binding cleft, which enables a lock-and-key type interaction with the lipid antigen. The structure provides a basis for the interaction between the highly conserved NKT TCR alpha-chain and the CD1d-antigen complex that is typified in innate immunity, and also indicates how variability of the NKT TCR beta-chain can impact on recognition of other CD1d-antigen complexes. These findings provide direct insight into how a T-cell receptor recognizes a lipid-antigen-presenting molecule of the immune system.
- The Protein Crystallography Unit, ARC Centre of Excellence in Structural and Functional Microbial Genomics, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: