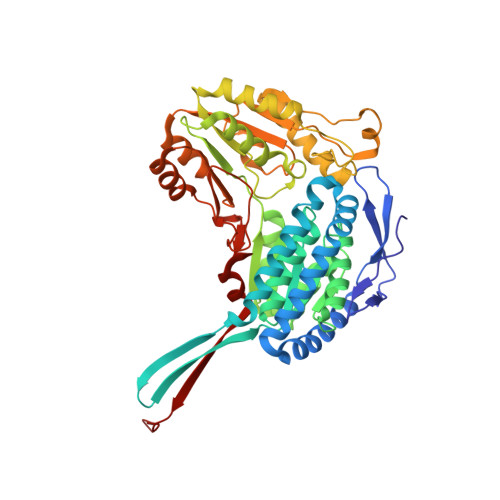
A Novel Cysteine-Nadph Covalent Adduct in Pseudomonas Aeruginosa Betaine Aldehyde Dehydrogenase Suggests Important Roles for the Reduced Nucleotide in the Reaction Mechanism
Diaz-Sanchez, A.G., Gonzalez-Segura, L., Rudino-Pinera, E., Lira-Rocha, A., Munoz-Clares, R.A.To be published.