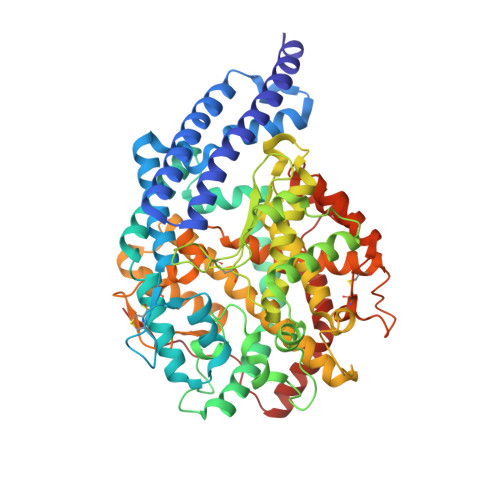
Crystal Structure of a Phosphonotripeptide K-26 in Complex with Angiotensin Converting Enzyme Homologue (Ance) from Drosophila Melanogaster.
Akif, M., Ntai, I., Sturrock, E.D., Isaac, R.E., Bachmann, B.O., Acharya, K.R.(2010) Biochem Biophys Res Commun 398: 532
- PubMed: 20599761
- DOI: https://doi.org/10.1016/j.bbrc.2010.06.113
- Primary Citation of Related Structures:
2XHM - PubMed Abstract:
Angiotensin-I converting enzyme (ACE, a zinc dependent dipeptidyl carboxypeptidase) is a major target of drugs due to its role in the modulation of blood pressure and cardiovascular disorders. Here we present a crystal structure of AnCE (an ACE homologue from Drosophila melanogaster with a single enzymatic domain) in complex with a natural product-phosphonotripeptide, K-26 at 1.96A resolution. The inhibitor binds exclusively in the S(1) and S(2) binding pockets of AnCE (coordinating the zinc ion) through ionic and hydrogen bond interactions. A detailed structural comparison of AnCE.K-26 complex with individual domains of human somatic ACE provides useful information for further exploration of ACE inhibitor pharmacophores involving phosphonic acids.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, UK.
Organizational Affiliation: