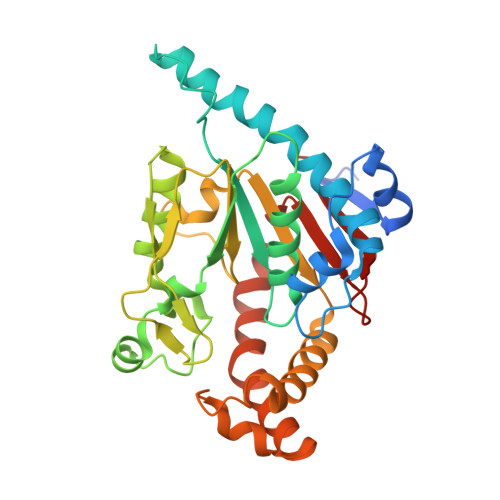
Location and conformation of pantothenate and its derivatives in Mycobacterium tuberculosis pantothenate kinase: insights into enzyme action
Chetnani, B., Kumar, P., Abhinav, K.V., Chhibber, M., Surolia, A., Vijayan, M.(2011) Acta Crystallogr D Biol Crystallogr 67: 774-783
- PubMed: 21904030 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444911024462
- Primary Citation Related Structures:
3AVO, 3AVP, 3AVQ - PubMed Abstract:
Previous studies of complexes of Mycobacterium tuberculosis PanK (MtPanK) with nucleotide diphosphates and nonhydrolysable analogues of nucleoside triphosphates in the presence or the absence of pantothenate established that the enzyme has dual specificity for ATP and GTP, revealed the unusual movement of ligands during enzyme action and provided information on the effect of pantothenate on the location and conformation of the nucleotides at the beginning and the end of enzyme action. The X-ray analyses of the binary complexes of MtPanK with pantothenate, pantothenol and N-nonylpantothenamide reported here demonstrate that in the absence of nucleotide these ligands occupy, with a somewhat open conformation, a location similar to that occupied by phosphopantothenate in the `end' complexes, which differs distinctly from the location of pantothenate in the closed conformation in the ternary `initiation' complexes. The conformation and the location of the nucleotide were also different in the initiation and end complexes. An invariant arginine appears to play a critical role in the movement of ligands that takes place during enzyme action. The work presented here completes the description of the locations and conformations of nucleoside diphosphates and triphosphates and pantothenate in different binary and ternary complexes, and suggests a structural rationale for the movement of ligands during enzyme action. The present investigation also suggests that N-alkylpantothenamides could be phosphorylated by the enzyme in the same manner as pantothenate.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore.
Organizational Affiliation: