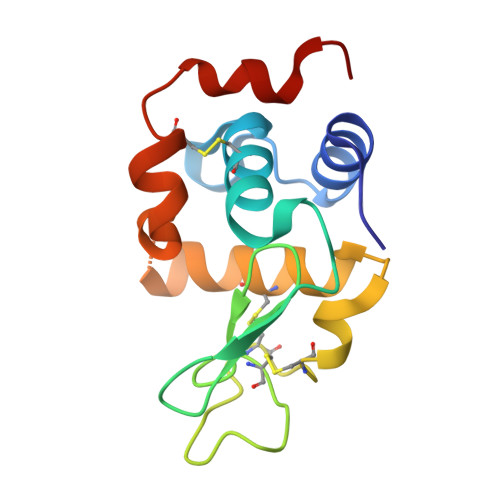
Structure of a lamprey variable lymphocyte receptor in complex with a protein antigen.
Velikovsky, C.A., Deng, L., Tasumi, S., Iyer, L.M., Kerzic, M.C., Aravind, L., Pancer, Z., Mariuzza, R.A.(2009) Nat Struct Mol Biol 16: 725-730
- PubMed: 19543291
- DOI: https://doi.org/10.1038/nsmb.1619
- Primary Citation of Related Structures:
3G39, 3G3A, 3G3B - PubMed Abstract:
Variable lymphocyte receptors (VLRs) are leucine-rich repeat proteins that mediate adaptive immunity in jawless vertebrates. VLRs are fundamentally different from the antibodies of jawed vertebrates, which consist of immunoglobulin (Ig) domains. We determined the structure of an anti-hen egg white lysozyme (HEL) VLR, isolated by yeast display, bound to HEL. The VLR, whose affinity resembles that of IgM antibodies, uses nearly all its concave surface to bind the protein, in addition to a loop that penetrates into the enzyme active site. The VLR-HEL structure combined with sequence analysis revealed an almost perfect match between ligand-contacting positions and positions with highest sequence diversity. Thus, it is likely that we have defined the generalized antigen-binding site of VLRs. We further demonstrated that VLRs can be affinity-matured by 13-fold to affinities as high as those of IgG antibodies, making VLRs potential alternatives to antibodies for biotechnology applications.
Organizational Affiliation:
Center for Advanced Research in Biotechnology, WM Keck Laboratory for Structural Biology, University of Maryland Biotechnology Institute, Rockville, Maryland, USA.