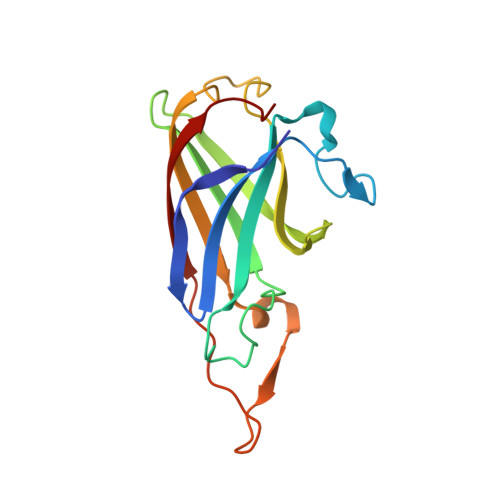
Structure and Function of P19, a High-Affinity Iron Transporter of the Human Pathogen Campylobacter jejuni.
Chan, A.C., Doukov, T.I., Scofield, M., Tom-Yew, S.A., Ramin, A.B., Mackichan, J.K., Gaynor, E.C., Murphy, M.E.(2010) J Mol Biology 401: 590-604
- PubMed: 20600116 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.06.038
- Primary Citation Related Structures:
3LZL, 3LZN, 3LZO, 3LZP, 3LZQ, 3LZR - PubMed Abstract:
Campylobacter jejuni, a major cause of acute bacterial diarrhea in humans, expresses numerous proteins to import diverse forms of essential iron. The expression of p19 and an adjacent iron transporter homologue (ftr1) is strongly induced upon iron limitation, suggesting a function in iron acquisition. Here, we show that the loss of P19 alone is detrimental to growth on iron-restricted media. Furthermore, metal binding analysis demonstrates that recombinant P19 has distinct copper and iron binding sites. Crystal structures of P19 have been solved to 1.41 A resolution, revealing an immunoglobulin-like fold. A P19 homodimer in which both monomers contribute ligands to two equivalent copper sites located adjacent to methionine-rich patches is observed. Copper coordination occurs via three histidine residues (His42, His95, and His132) and Met88. A solvent channel lined with conserved acidic residues leads to the copper site. Soaking crystals with a solution of manganese as iron analog reveals a second metal binding site in this solvent channel (metal-metal distance, 7.7 A). Glu44 lies between the metal sites and displays multiple conformations in the crystal structures, suggesting a role in regulating metal-metal interaction. Dimerization is shown to be metal dependent in vitro and is detected in vivo by cross-linking.
- Department of Microbiology and Immunology, Life Sciences Institute, 2350 Health Sciences Mall, The University of British Columbia, Vancouver, BC, Canada V6T 1Z3.
Organizational Affiliation: