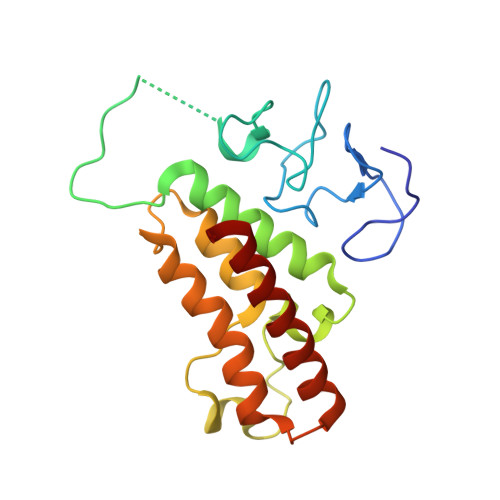
TRIM24 links a non-canonical histone signature to breast cancer.
Tsai, W.W., Wang, Z., Yiu, T.T., Akdemir, K.C., Xia, W., Winter, S., Tsai, C.Y., Shi, X., Schwarzer, D., Plunkett, W., Aronow, B., Gozani, O., Fischle, W., Hung, M.C., Patel, D.J., Barton, M.C.(2010) Nature 468: 927-932
- PubMed: 21164480
- DOI: https://doi.org/10.1038/nature09542
- Primary Citation Related Structures:
3O33, 3O34, 3O35, 3O36, 3O37 - PubMed Abstract:
Recognition of modified histone species by distinct structural domains within 'reader' proteins plays a critical role in the regulation of gene expression. Readers that simultaneously recognize histones with multiple marks allow transduction of complex chromatin modification patterns into specific biological outcomes. Here we report that chromatin regulator tripartite motif-containing 24 (TRIM24) functions in humans as a reader of dual histone marks by means of tandem plant homeodomain (PHD) and bromodomain (Bromo) regions. The three-dimensional structure of the PHD-Bromo region of TRIM24 revealed a single functional unit for combinatorial recognition of unmodified H3K4 (that is, histone H3 unmodified at lysine 4, H3K4me0) and acetylated H3K23 (histone H3 acetylated at lysine 23, H3K23ac) within the same histone tail. TRIM24 binds chromatin and oestrogen receptor to activate oestrogen-dependent genes associated with cellular proliferation and tumour development. Aberrant expression of TRIM24 negatively correlates with survival of breast cancer patients. The PHD-Bromo of TRIM24 provides a structural rationale for chromatin activation through a non-canonical histone signature, establishing a new route by which chromatin readers may influence cancer pathogenesis.
- Department of Biochemistry and Molecular Biology, Program in Genes and Development, Graduate School of Biomedical Sciences, University of Texas M.D. Anderson Cancer Center, Houston, Texas 77030, USA.
Organizational Affiliation: