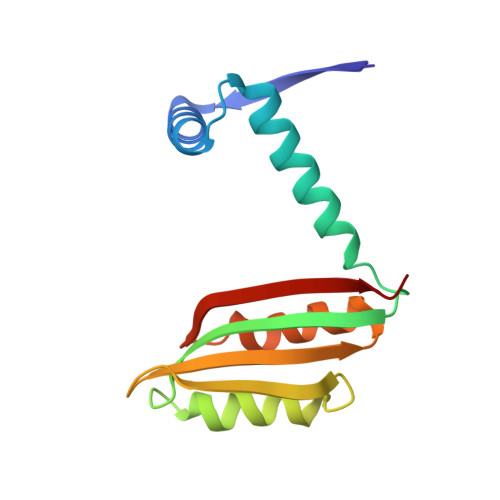
Structural basis of low-affinity nickel binding to the nickel-responsive transcription factor NikR from Escherichia coli.
Phillips, C.M., Schreiter, E.R., Stultz, C.M., Drennan, C.L.(2010) Biochemistry 49: 7830-7838
- PubMed: 20704276 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi100923j
- Primary Citation Related Structures:
3OD2 - PubMed Abstract:
Escherichia coli NikR regulates cellular nickel uptake by binding to the nik operon in the presence of nickel and blocking transcription of genes encoding the nickel uptake transporter. NikR has two binding affinities for the nik operon: a nanomolar dissociation constant with stoichiometric nickel and a picomolar dissociation constant with excess nickel [Bloom, S. L., and Zamble, D. B. (2004) Biochemistry 43, 10029-10038; Chivers, P. T., and Sauer, R. T. (2002) Chem. Biol. 9, 1141-1148]. While it is known that the stoichiometric nickel ions bind at the NikR tetrameric interface [Schreiter, E. R., et al. (2003) Nat. Struct. Biol. 10, 794-799; Schreiter, E. R., et al. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 13676-13681], the binding sites for excess nickel ions have not been fully described. Here we have determined the crystal structure of NikR in the presence of excess nickel to 2.6 A resolution and have obtained nickel anomalous data (1.4845 A) in the presence of excess nickel for both NikR alone and NikR cocrystallized with a 30-nucleotide piece of double-stranded DNA containing the nik operon. These anomalous data show that excess nickel ions do not bind to a single location on NikR but instead reveal a total of 22 possible low-affinity nickel sites on the NikR tetramer. These sites, for which there are six different types, are all on the surface of NikR, and most are found in both the NikR alone and NikR-DNA structures. Using a combination of crystallographic data and molecular dynamics simulations, the nickel sites can be described as preferring octahedral geometry, utilizing one to three protein ligands (typically histidine) and at least two water molecules.
- Department of Chemistry, Massachusetts Institute of Technology,Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: