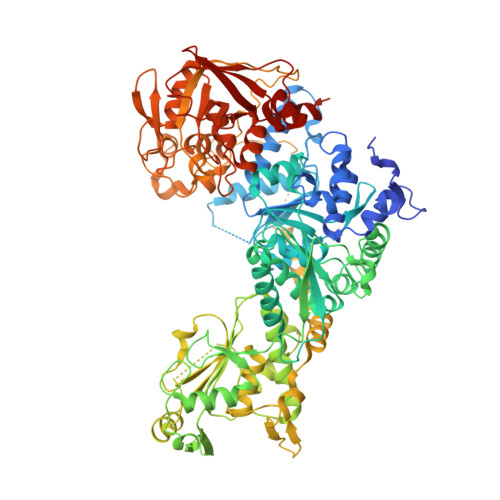
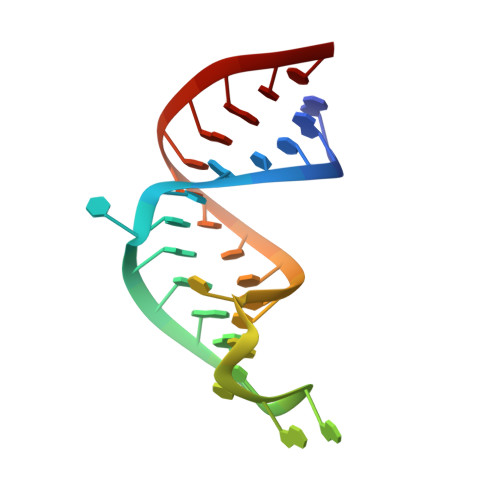
Accommodating variety in iron-responsive elements: Crystal structure of transferrin receptor 1 B IRE bound to iron regulatory protein 1.
Walden, W.E., Selezneva, A., Volz, K.(2012) FEBS Lett 586: 32-35
- PubMed: 22119729 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2011.11.018
- Primary Citation Related Structures:
3SN2 - PubMed Abstract:
Iron responsive elements (IREs) are short stem-loop structures found in several mRNAs encoding proteins involved in cellular iron metabolism. Iron regulatory proteins (IRPs) control iron homeostasis through differential binding to the IREs, accommodating any sequence or structural variations that the IREs may present. Here we report the structure of IRP1 in complex with transferrin receptor 1 B (TfR B) IRE, and compare it to the complex with ferritin H (Ftn H) IRE. The two IREs are bound to IRP1 through nearly identical protein-RNA contacts, although their stem conformations are significantly different. These results support the view that binding of different IREs with IRP1 depends both on protein and RNA conformational plasticity, adapting to RNA variation while retaining conserved protein-RNA contacts.
- Department of Microbiology and Immunology, University of Illinois at Chicago, Chicago, IL 60612, USA.
Organizational Affiliation: