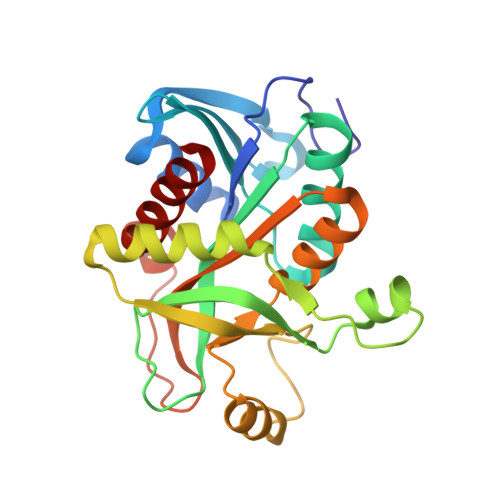
New phosphate binding sites in the crystal structure of Escherichia coli purine nucleoside phosphorylase complexed with phosphate and formycin A.
Stefanic, Z., Narczyk, M., Mikleusevic, G., Wielgus-Kutrowska, B., Bzowska, A., Luic, M.(2012) FEBS Lett 586: 967-971
- PubMed: 22569248 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2012.02.039
- Primary Citation Related Structures:
3UT6 - PubMed Abstract:
Purine nucleoside phosphorylase (PNP) from Escherichia coli is a homohexamer that catalyses the phosphorolytic cleavage of the glycosidic bond of purine nucleosides. The first crystal structure of the ternary complex of this enzyme (with a phosphate ion and formycin A), which is biased by neither the presence of an inhibitor nor sulfate as a precipitant, is presented. The structure reveals, in some active sites, an unexpected and never before observed binding site for phosphate and exhibits a stoichiometry of two phosphate molecules per enzyme subunit. Moreover, in these active sites, the phosphate and nucleoside molecules are found not to be in direct contact. Rather, they are bridged by three water molecules that occupy the "standard" phosphate binding site.
- Rudjer Bošković Institute, Bijenička 54, 10000 Zagreb, Croatia.
Organizational Affiliation: