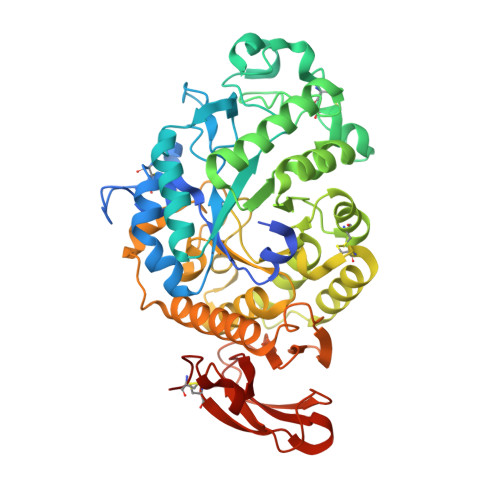
A Novel Multifunctional alpha-Amylase from the Thermophilic Fungus Malbranchea cinnamomea: Biochemical Characterization and Three-Dimensional Structure.
Han, P., Zhou, P., Hu, S., Yang, S., Yan, Q., Jiang, Z.Q.(2013) Appl Biochem Biotechnol 170: 420-435
- PubMed: 23536251
- DOI: https://doi.org/10.1007/s12010-013-0198-y
- Primary Citation of Related Structures:
3VM7 - PubMed Abstract:
A novel α-amylase (McAmyA) from the thermophilic fungus, Malbranchea cinnamomea was purified, characterized and crystallized in the present study. McAmyA was purified to apparent homogeneity with a molecular mass of 60.3 kDa on SDS-PAGE. The enzyme exhibited maximal activity at pH 6.5 and was stable within pH 5.0-10.0. It was most active at 65 °C and was stable up to 50 °C. McAmyA was capable of hydrolyzing amylose, starch, amylopectin, pullulan, cyclodextrins and maltooligosaccharides. The full-length cDNA of an α-amylase gene (McAmyA) from the strain was cloned. McAmyA consisted of a 1,476-bp open reading frame encoding 492 amino acids. It displayed the highest amino acid sequence homology (less than 60 %) with the reported α-amylases. The crystal structure of McAmyA was solved at a resolution of 2.25 Å (PDB code 3VM7). The overall structure of McAmyA reveals three domains with ten α helices and 14 β strands, and the putative catalytic residues are positioned at domain A with somewhat different secondary structural circumstances compared with typical α-amylases.
- Department of Biotechnology, College of Food Science and Nutritional Engineering, China Agricultural University, PO Box 294, No.17 Qinghua Donglu, Haidian District, Beijing, 100083, China.
Organizational Affiliation: