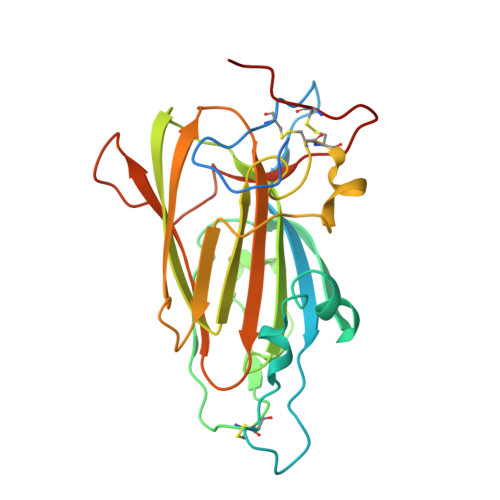
Structural Basis for Promiscuity and Specificity During Candida Glabrata Invasion of Host Epithelia.
Maestre-Reyna, M., Diderrich, R., Veelders, M.S., Eulenburg, G., Kalugin, V., Bruckner, S., Keller, P., Rupp, S., Mosch, H., Essen, L.-O.(2012) Proc Natl Acad Sci U S A 109: 16864
- PubMed: 23035251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1207653109
- Primary Citation Related Structures:
4AF9, 4AFA, 4AFB, 4AFC, 4ASL - PubMed Abstract:
The human pathogenic yeast Candida glabrata harbors more than 20 surface-exposed, epithelial adhesins (Epas) for host cell adhesion. The Epa family recognizes host glycans and discriminates between target tissues by their adhesin (A) domains, but a detailed structural basis for ligand-binding specificity of Epa proteins has been lacking so far. In this study, we provide high-resolution crystal structures of the Epa1A domain in complex with different carbohydrate ligands that reveal how host cell mucin-type O-glycans are recognized and allow a structure-guided classification of the Epa family into specific subtypes. Further detailed structural and functional characterization of subtype-switched Epa1 variants shows that specificity is governed by two inner loops, CBL1 and CBL2, involved in calcium binding as well as by three outer loops, L1, L2, and L3. In summary, our study provides the structural basis for promiscuity and specificity of Epa adhesins, which might further contribute to developing anti-adhesive antimycotics and combating Candida colonization.
- Biomedical Research Center/Department of Chemistry, Philipps-Universität Marburg, D-35032 Marburg, Germany.
Organizational Affiliation: