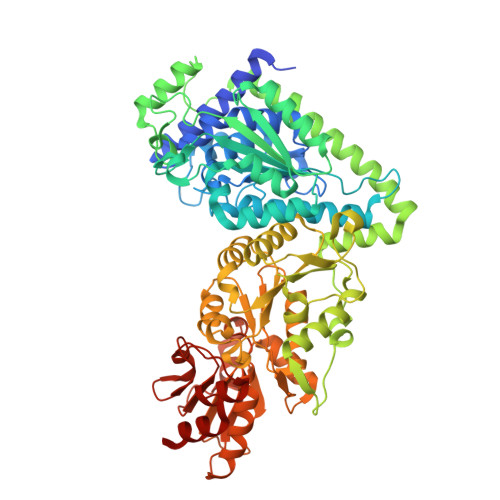
High Resolution Crystal Structures of Lactobacillus Salivarius Transketolase in the Presence and Absence of Thiamine Pyrophosphate
Lukacik, P., Lobley, C.M.C., Bumann, M., Arena De Souza, V., Owens, R.J., O'Toole, P.W., Walsh, M.A.(2015) Acta Crystallogr Sect F Struct Biol Cryst Commun 71: 1327
- PubMed: 26457526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X1501657X
- Primary Citation Related Structures:
4C7V, 4C7X - PubMed Abstract:
Probiotic bacterial strains have been shown to enhance the health of the host through a range of mechanisms including colonization, resistance against pathogens, secretion of antimicrobial compounds and modulation of the activity of the innate immune system. Lactobacillus salivarius UCC118 is a well characterized probiotic strain which survives intestinal transit and has many desirable host-interaction properties. Probiotic bacteria display a wide range of catabolic activities, which determine their competitiveness in vivo. Some lactobacilli are heterofermentative and can metabolize pentoses, using a pathway in which transketolase and transaldolase are key enzymes. L. salivarius UCC118 is capable of pentose utilization because it encodes the key enzymes on a megaplasmid. The crystal structures of the megaplasmid-encoded transketolase with and without the enzyme cofactor thiamine pyrophosphate have been determined. Comparisons with other known transketolase structures reveal a high degree of structural conservation in both the catalytic site and the overall conformation. This work extends structural knowledge of the transketolases to the industrially and commercially important Lactobacillus genus.
- Diamond Light Source, Harwell Science and Innovation Campus, Didcot OX11 0DE, England.
Organizational Affiliation: