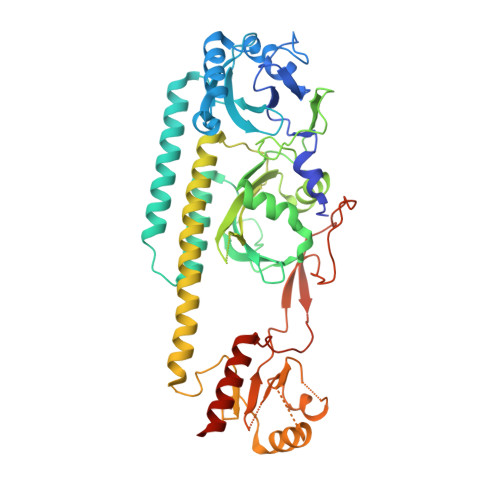
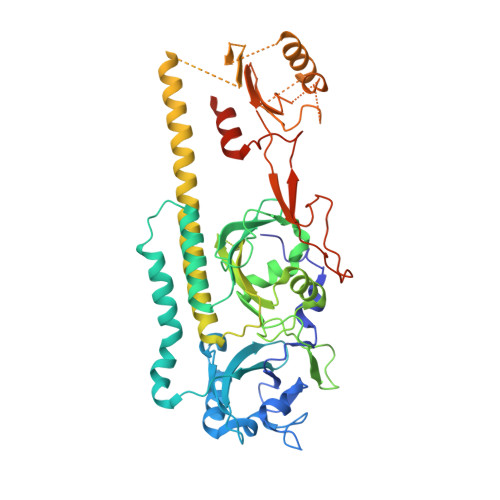
Crystal structure of the photosensing module from a red/far-red light-absorbing plant phytochrome.
Burgie, E.S., Bussell, A.N., Walker, J.M., Dubiel, K., Vierstra, R.D.(2014) Proc Natl Acad Sci U S A 111: 10179-10184
- PubMed: 24982198 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1403096111
- Primary Citation Related Structures:
4OUR - PubMed Abstract:
Many aspects of plant photomorphogenesis are controlled by the phytochrome (Phy) family of bilin-containing photoreceptors that detect red and far-red light by photointerconversion between a dark-adapted Pr state and a photoactivated Pfr state. Whereas 3D models of prokaryotic Phys are available, models of their plant counterparts have remained elusive. Here, we present the crystal structure of the photosensing module (PSM) from a seed plant Phy in the Pr state using the PhyB isoform from Arabidopsis thaliana. The PhyB PSM crystallized as a head-to-head dimer with strong structural homology to its bacterial relatives, including a 5(Z)syn, 10(Z)syn, 15(Z)anti configuration of the phytochromobilin chromophore buried within the cGMP phosphodiesterase/adenylyl cyclase/FhlA (GAF) domain, and a well-ordered hairpin protruding from the Phy-specific domain toward the bilin pocket. However, its Per/Arnt/Sim (PAS) domain, knot region, and helical spine show distinct structural differences potentially important to signaling. Included is an elongated helical spine, an extended β-sheet connecting the GAF domain and hairpin stem, and unique interactions between the region upstream of the PAS domain knot and the bilin A and B pyrrole rings. Comparisons of this structure with those from bacterial Phys combined with mutagenic studies support a toggle model for photoconversion that engages multiple features within the PSM to stabilize the Pr and Pfr end states after rotation of the D pyrrole ring. Taken together, this Arabidopsis PhyB structure should enable molecular insights into plant Phy signaling and provide an essential scaffold to redesign their activities for agricultural benefit and as optogenetic reagents.
- Department of Genetics, University of Wisconsin-Madison, Madison, WI 53706.
Organizational Affiliation: