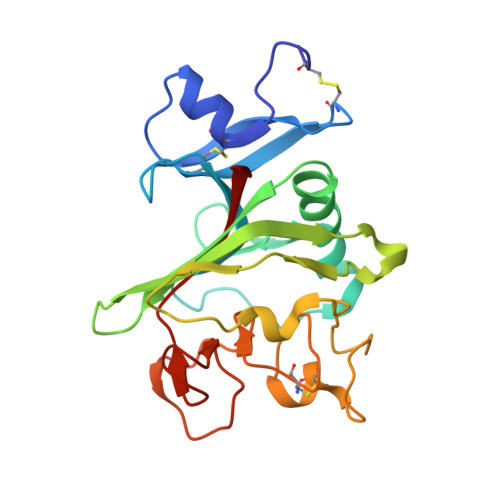
Human ficolin-2 recognition versatility extended: An update on the binding of ficolin-2 to sulfated/phosphated carbohydrates.
Laffly, E., Lacroix, M., Martin, L., Vassal-Stermann, E., Thielens, N.M., Gaboriaud, C.(2014) FEBS Lett 588: 4694-4700
- PubMed: 25447524
- DOI: https://doi.org/10.1016/j.febslet.2014.10.042
- Primary Citation of Related Structures:
4R9J, 4R9T - PubMed Abstract:
Ficolin-2 has been reported to bind to DNA and heparin, but the mechanism involved has not been thoroughly investigated. X-ray studies of the ficolin-2 fibrinogen-like domain in complex with several new ligands now show that sulfate and phosphate groups are prone to bind to the S3 binding site of the protein. Composed of Arg132, Asp133, Thr136 and Lys221, the S3 site was previously shown to mainly bind N-acetyl groups. Furthermore, DNA and heparin compete for binding to ficolin-2. Mutagenesis studies reveal that Arg132, and to a lesser extent Asp133, are important for this binding property. The versatility of the S3 site in binding N-acetyl, sulfate and phosphate groups is discussed through comparisons with homologous fibrinogen-like recognition proteins.
- Univ. Grenoble Alpes, IBS, F-38044 Grenoble, France; CNRS, IBS, F-38044 Grenoble, France; CEA, IBS, F-38044 Grenoble, France.
Organizational Affiliation: