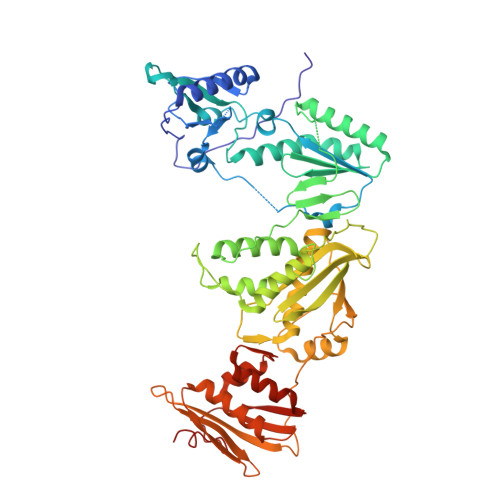
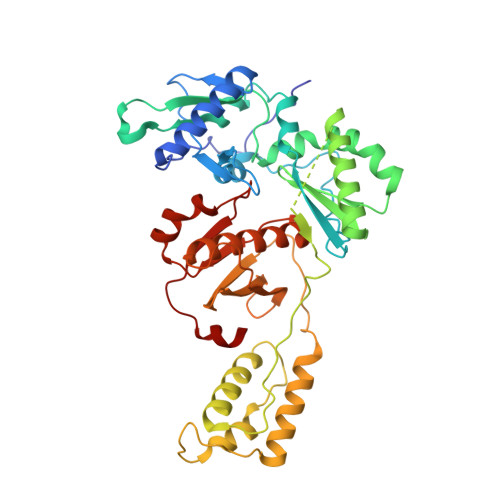
Structure-Based Evaluation of Non-nucleoside Inhibitors with Improved Potency and Solubility That Target HIV Reverse Transcriptase Variants.
Frey, K.M., Puleo, D.E., Spasov, K.A., Bollini, M., Jorgensen, W.L., Anderson, K.S.(2015) J Med Chem 58: 2737-2745
- PubMed: 25700160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm501908a
- Primary Citation Related Structures:
4RW4, 4RW6, 4RW7, 4RW8, 4RW9 - PubMed Abstract:
The development of novel non-nucleoside inhibitors (NNRTIs) with activity against variants of HIV reverse transcriptase (RT) is crucial for overcoming treatment failure. The NNRTIs bind in an allosteric pocket in RT ∼10 Å away from the active site. Earlier analogues of the catechol diether compound series have picomolar activity against HIV strains with wild-type RT but lose potency against variants with single Y181C and double K103N/Y181C mutations. As guided by structure-based and computational studies, removal of the 5-Cl substitution of compound 1 on the catechol aryl ring system led to a new analogue compound 2 that maintains greater potency against Y181C and K103N/Y181C variants and better solubility (510 μg/mL). Crystal structures were determined for wild-type, Y181C, and K103N/Y181C RT in complex with both compounds 1 and 2 to understand the structural basis for these findings. Comparison of the structures reveals that the Y181C mutation destabilizes the binding mode of compound 1 and disrupts the interactions with residues in the pocket. Compound 2 maintains the same conformation in wild-type and mutant structures, in addition to several interactions with the NNRTI binding pocket. Comparison of the six crystal structures will assist in the understanding of compound binding modes and future optimization of the catechol diether series.
- †Department of Pharmacology, ‡Department of Chemistry, Yale University, New Haven, Connecticut 06520-8066, United States.
Organizational Affiliation: