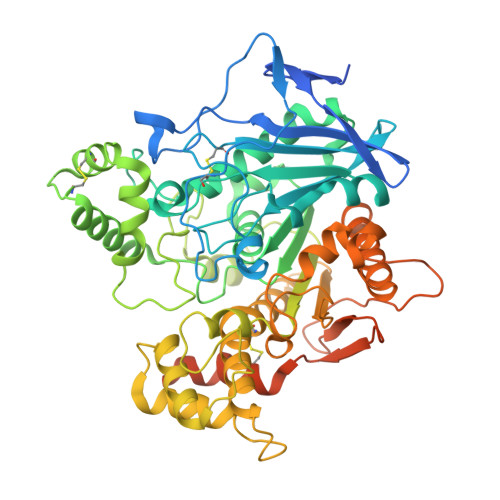
Discovery, biological evaluation, and crystal structure of a novel nanomolar selective butyrylcholinesterase inhibitor.
Brus, B., Kosak, U., Turk, S., Pislar, A., Coquelle, N., Kos, J., Stojan, J., Colletier, J.P., Gobec, S.(2014) J Med Chem 57: 8167-8179
- PubMed: 25226236 Search on PubMed
- DOI: https://doi.org/10.1021/jm501195e
- Primary Citation Related Structures:
4TPK - PubMed Abstract:
Butyrylcholinesterase (BChE) is regarded as a promising drug target as its levels and activity significantly increase in the late stages of Alzheimer's disease. To discover novel BChE inhibitors, we used a hierarchical virtual screening protocol followed by biochemical evaluation of 40 highest scoring hit compounds. Three of the compounds identified showed significant inhibitory activities against BChE. The most potent, compound 1 (IC50 = 21.3 nM), was resynthesized and resolved into its pure enantiomers. A high degree of stereoselective activity was revealed, and a dissociation constant of 2.7 nM was determined for the most potent stereoisomer (+)-1. The crystal structure of human BChE in complex with compound (+)-1 was solved, revealing the binding mode and providing clues for potential optimization. Additionally, compound 1 inhibited amyloid β(1-42) peptide self-induced aggregation into fibrils (by 61.7% at 10 μM) and protected cultured SH-SY5Y cells against amyloid-β-induced toxicity. These data suggest that compound 1 represents a promising candidate for hit-to-lead follow-up in the drug-discovery process against Alzheimer's disease.
- Faculty of Pharmacy, University of Ljubljana , Aškerčeva 7, 1000 Ljubljana, Slovenia.
Organizational Affiliation: