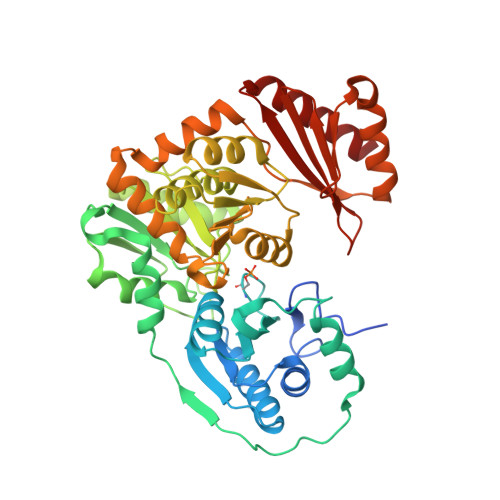
Structural and functional characterization of the phosphoglucomutase from Xanthomonas citri subsp. citri.
Goto, L.S., Vessoni Alexandrino, A., Malvessi Pereira, C., Silva Martins, C., D'Muniz Pereira, H., Brandao-Neto, J., Marques Novo-Mansur, M.T.(2016) Biochim Biophys Acta 1864: 1658-1666
- PubMed: 27567706 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2016.08.014
- Primary Citation Related Structures:
5BMN, 5BMP, 5KL0 - PubMed Abstract:
Citrus canker, caused by bacteria Xanthomonas citri subsp. citri, can affect all economically important varieties of citrus. Studying Xanthomonas genes related to the invasive capacity may improve the knowledge on how this works and ultimately use the information to avoid the disease. Some annotated genes from Xanthomonas citri subsp. citri published genome are addressed to an interesting class of genes named "pathogenicity, virulence and adaptation". One of them is xanA, which encodes a predicted phosphoglucomutase. Phosphoglucomutases are ubiquitous enzymes among the living kingdoms that play roles in carbohydrate metabolism, catalyzing the reversible conversion of 1- to 6-phosphoglucose. In Xanthomonas, phosphoglucomutase activity is required to synthesize precursors of the pathogenesis-related polysaccharide xanthan. In this work, a characterization of this gene product is presented by structural and functional studies. Molecular cloning was used for heterologous expression and deletion of xanA. A Michaelis-Menten kinetics model was obtained using the recombinant protein. The protein structure was also determined by X-ray diffraction on the recombinant enzyme substrate-free, bound to glucose-1,6-biphosphate and to glucose-1-phosphate. Deletion of xanA was done with a suicide plasmid construct and the obtained mutant was tested for pathogenic capacity. This study is the first describing the properties of the Xanthomonas citri subsp. citri phosphoglucomutase.
- Laboratório de Bioquímica e Biologia Molecular Aplicada - LBBMA, Departamento de Genética e Evolução, Universidade Federal de São Carlos, São Carlos, SP, Brazil. Electronic address: seiji_l@hotmail.com.
Organizational Affiliation: